Quick filters:
Page 1 of 1
Proto oncogene Stock Photos and Images
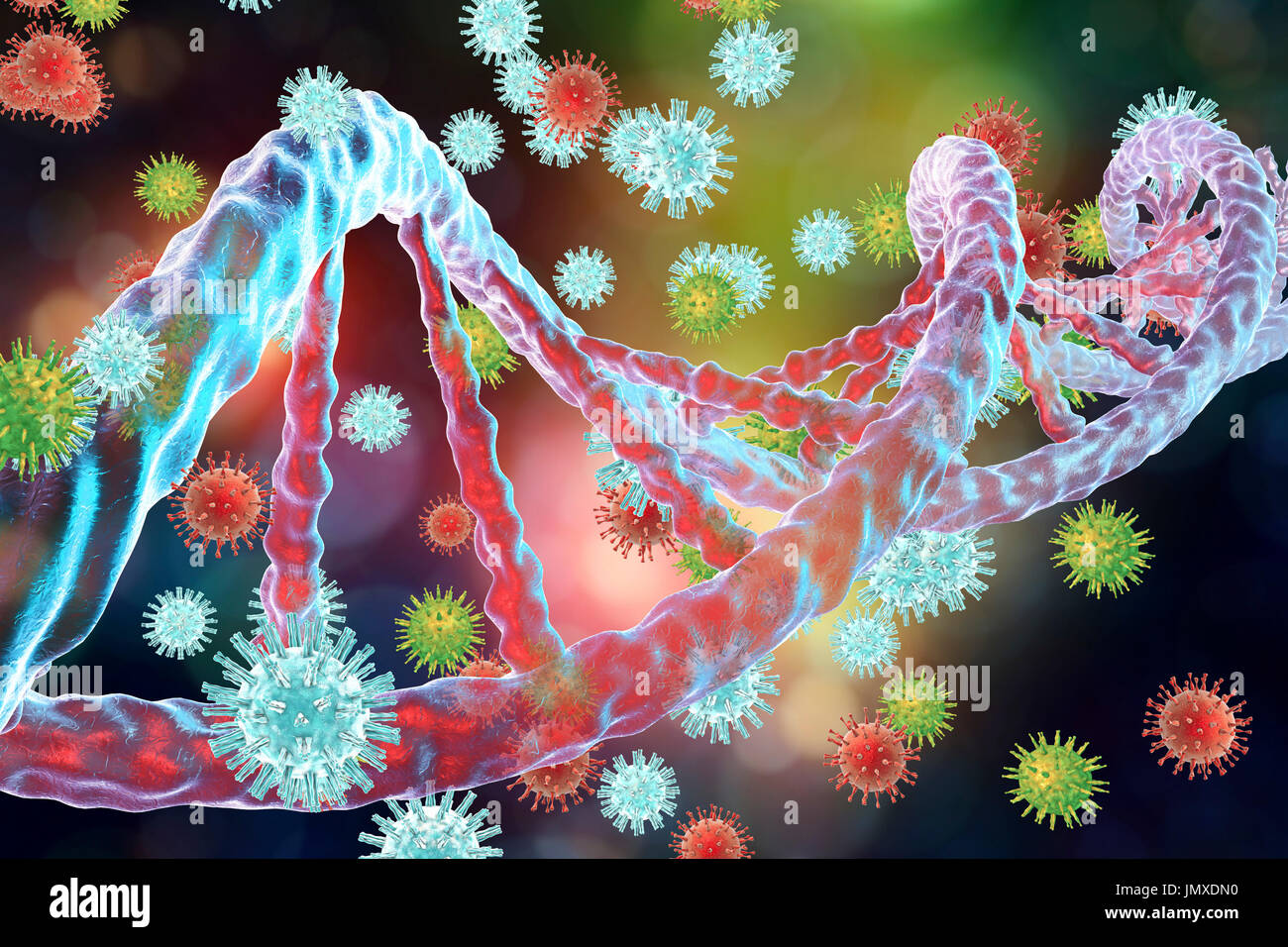
RFJMXDN0–Conceptual image for interaction between viruses and host-cell DNA (deoxyribonucleic acid). Integration of viruses into DNA is the key step in oncogenesis. Several viruses, such as hepatitis B virus, papillomavirus and other, can integrate into host DNA as insertional mutagens causing the activation of a cellular proto-oncogene which eventually leads to uncontrolled cell multiplication and cancer development.
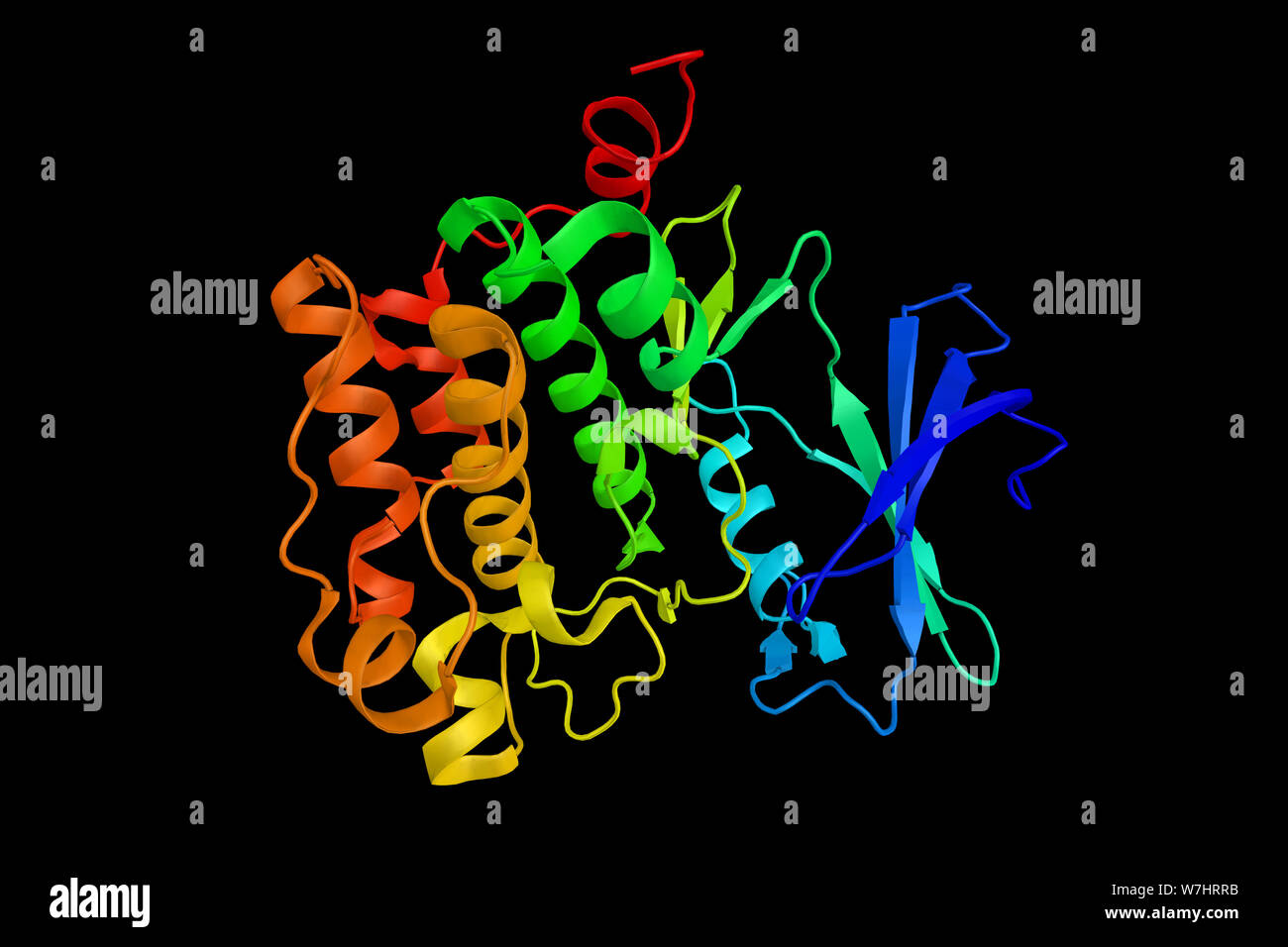
RFW7HRRB–PIM1, a proto-oncogene which encodes for the serine/threonine kinase of the same name. 3d rendering.
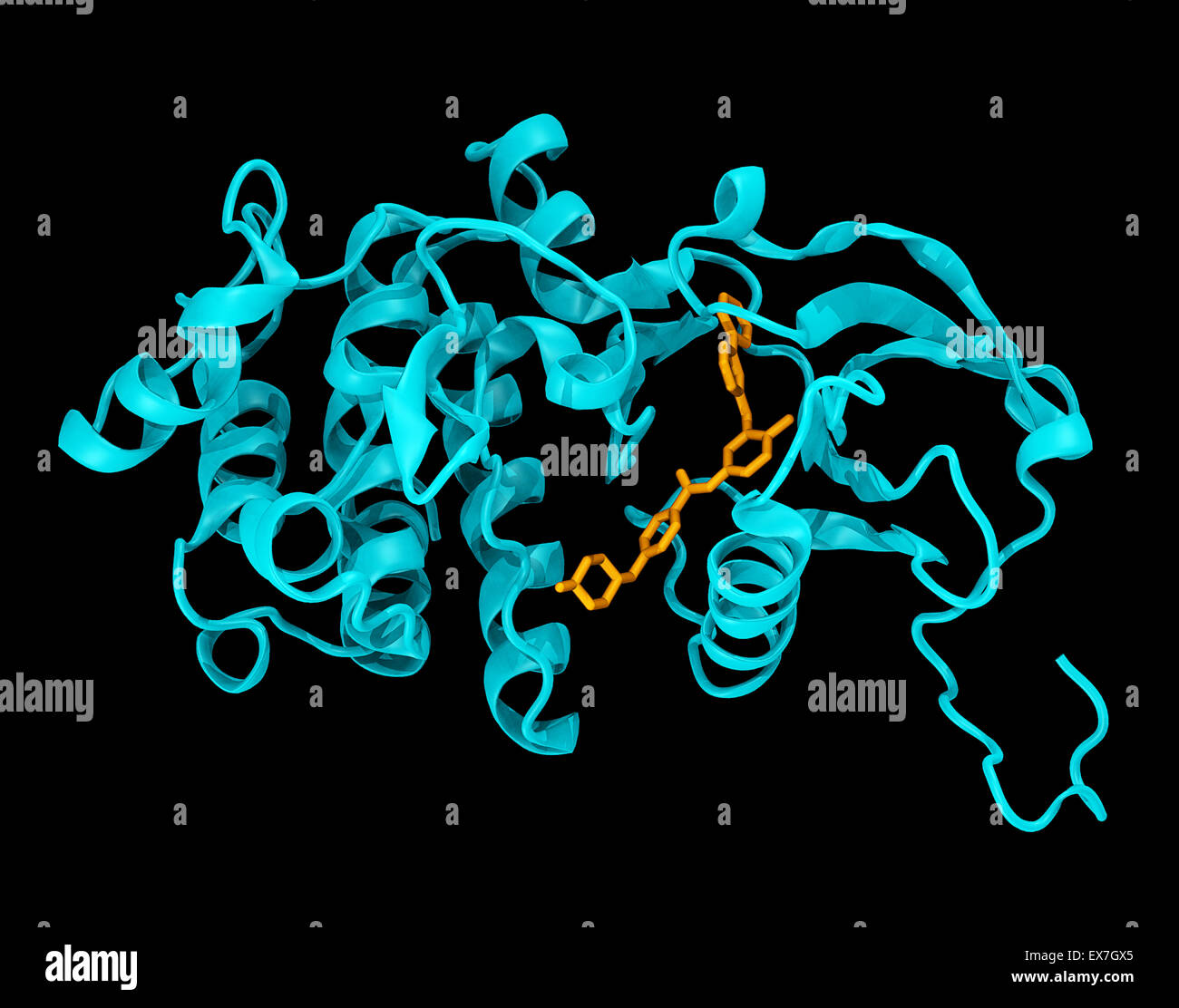
RMEX7GX5–Crystal structure of the c-abl kinase domain (cyan) in complex with imatinib (Gleevec) (colored orange).
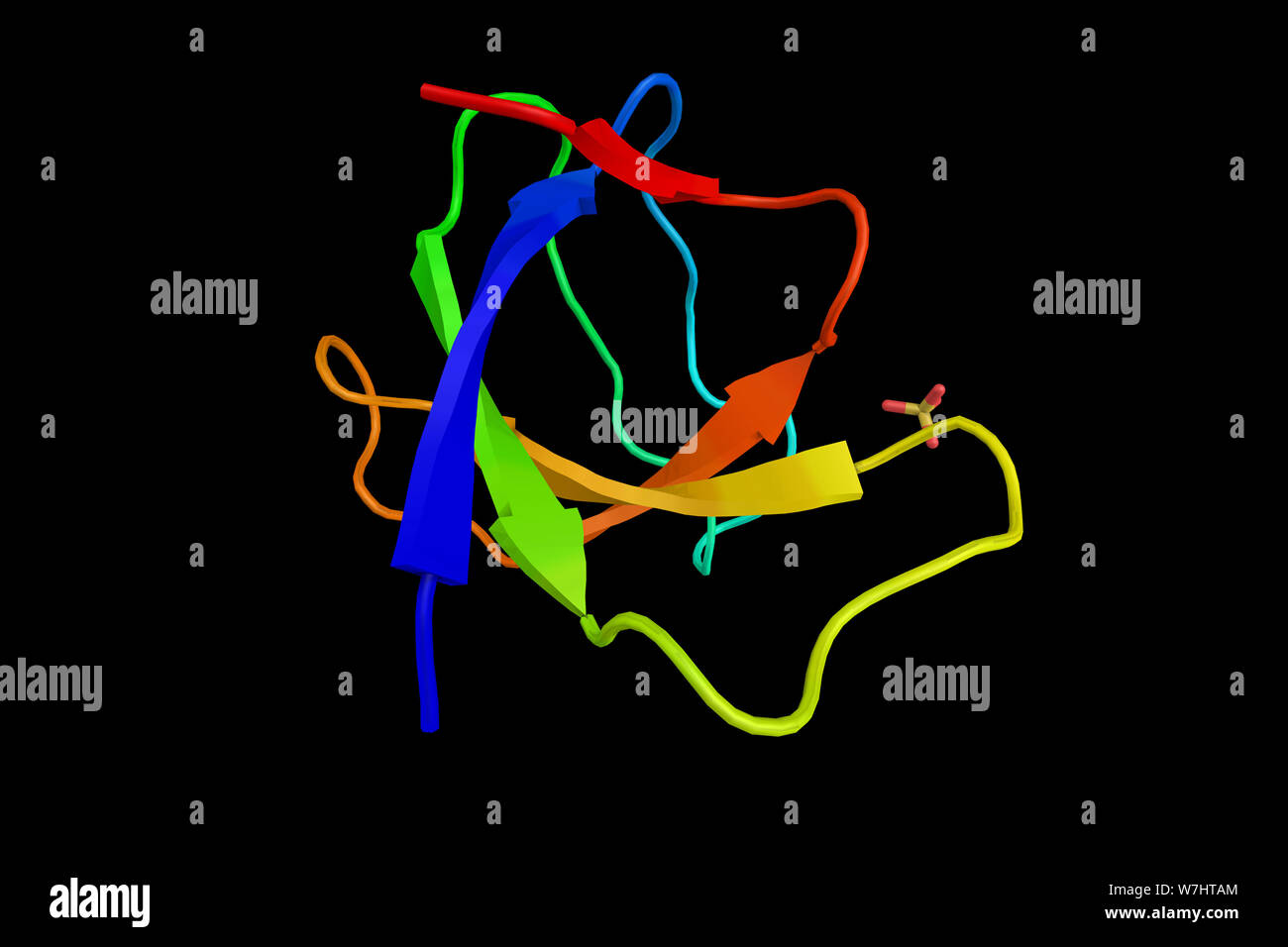
RFW7HTAM–Proto-oncogene tyrosine-protein kinase Yes, a protein which has tyrosine kinase activity and belongs to the src family of proteins. 3d rendering.
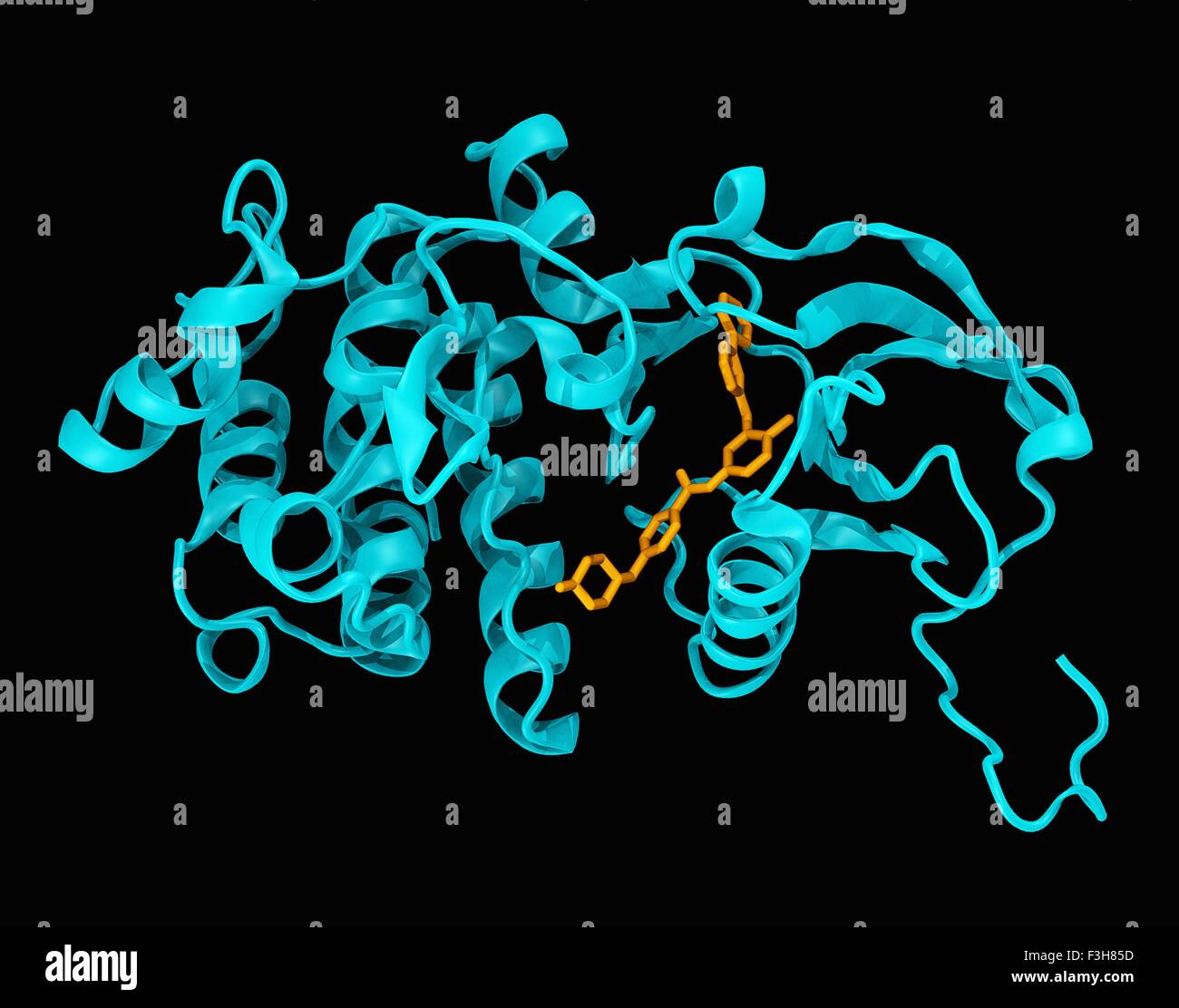
RFF3H85D–Crystal structure of the c-abl kinase domain (cyan) in complex with imatinib (Gleevec) (colored orange)
RF2R729BA–Conceptual image for interaction between viruses and host-cell DNA. Intergation of viruses into DNA is the key step in oncogenesis. Several viruses, s
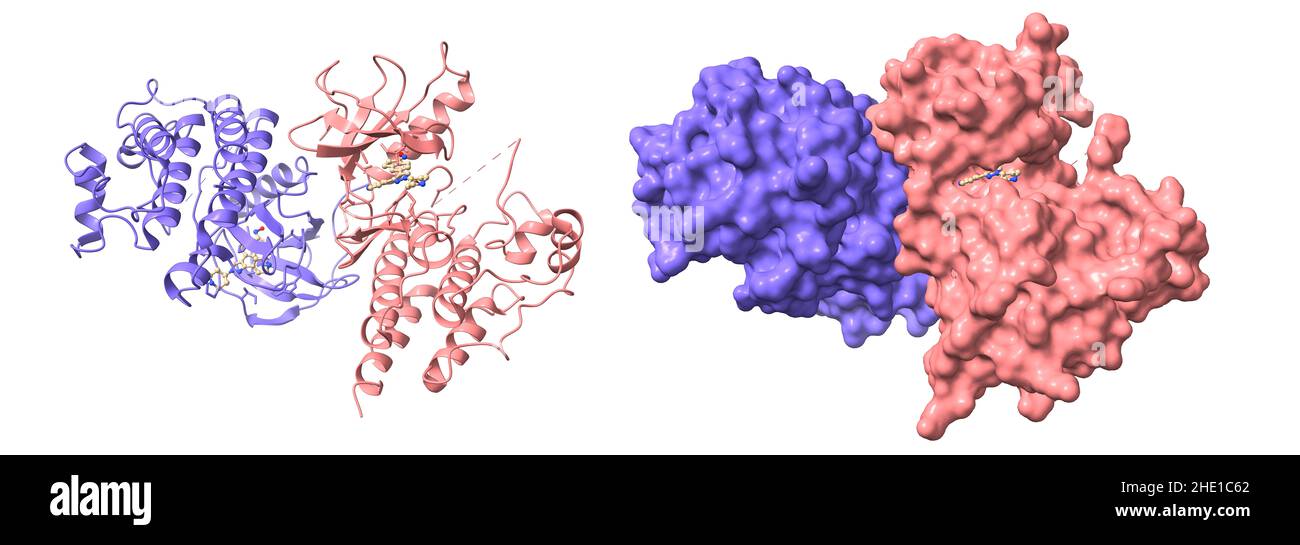
RF2HE1C62–Crystal structure of c-raf (raf-1), a mitogen-activated protein kinase dimer. 3D cartoon and Gaussian surface models, chain id color scheme, PDB 3omv
RFJMXDMY–Conceptual image for interaction between viruses and host-cell DNA (deoxyribonucleic acid). Integration of viruses into DNA is the key step in oncogenesis. Several viruses, such as hepatitis B virus, papillomavirus and other, can integrate into host DNA as insertional mutagens causing the activation of a cellular proto-oncogene which eventually leads to uncontrolled cell multiplication and cancer development.
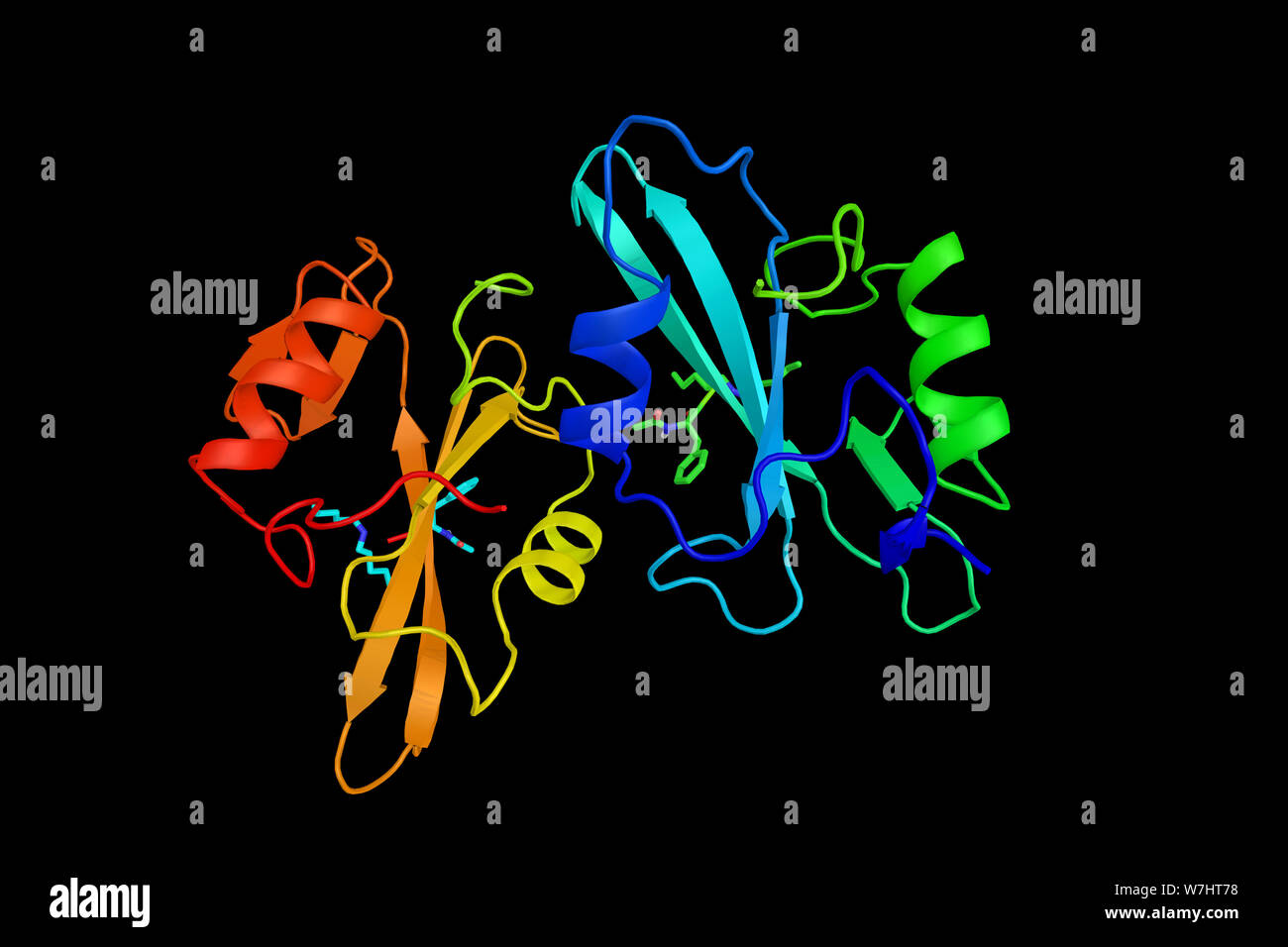
RFW7HT78–Proto-oncogene tyrosine-protein kinase Src, a non-receptor tyrosine kinase protein which phosphorylates specific tyrosine residues in other proteins.
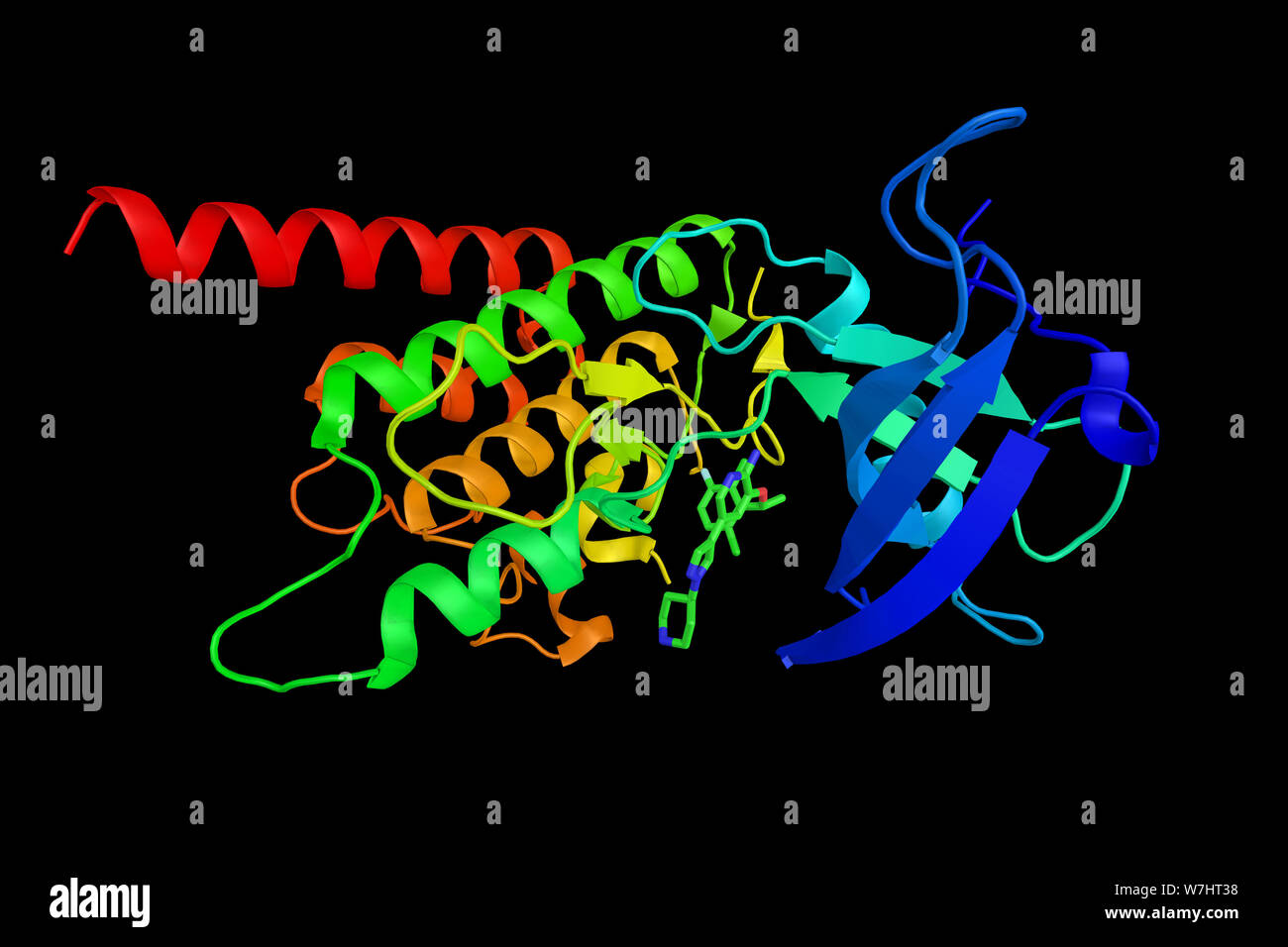
RFW7HT38–Proto-oncogene tyrosine-protein kinase ROS, an enzyme which is a proto-oncogene, highly expressed in a variety of tumor cell lines. 3d rendering.
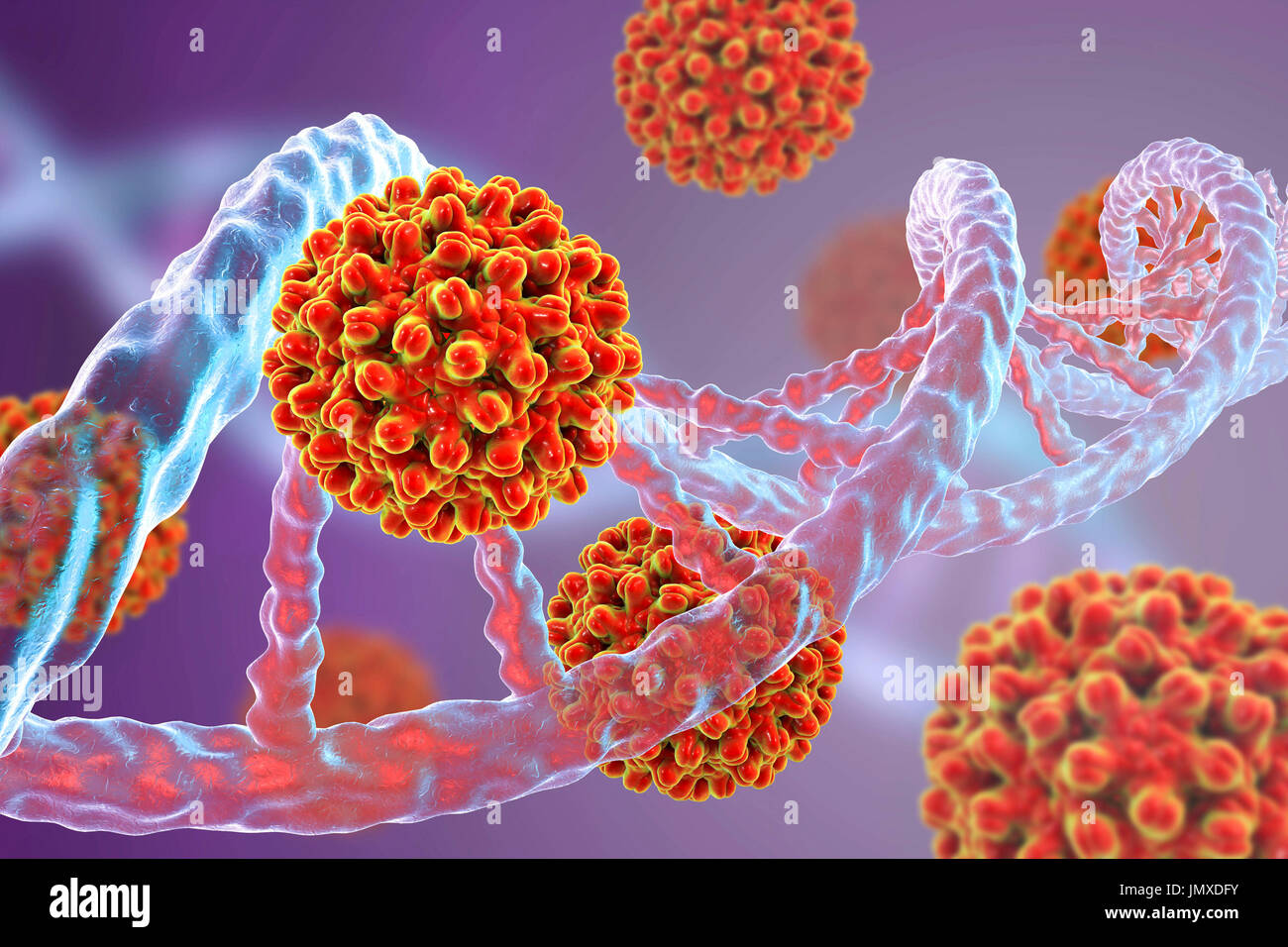
RFJMXDFY–Hepatitis B viruses and DNA. Conceptual image for viral oncogenesis. Hepatitis B viruses (HBV) can integrate into host DNA as insertional mutagens causing the activation of a cellular proto-oncogene. Integration of viral DNA into the human genome is considered an early event in the carcinogenic process and can induce, through insertional mutagenesis, the alteration of gene expression and chromosomal instability.
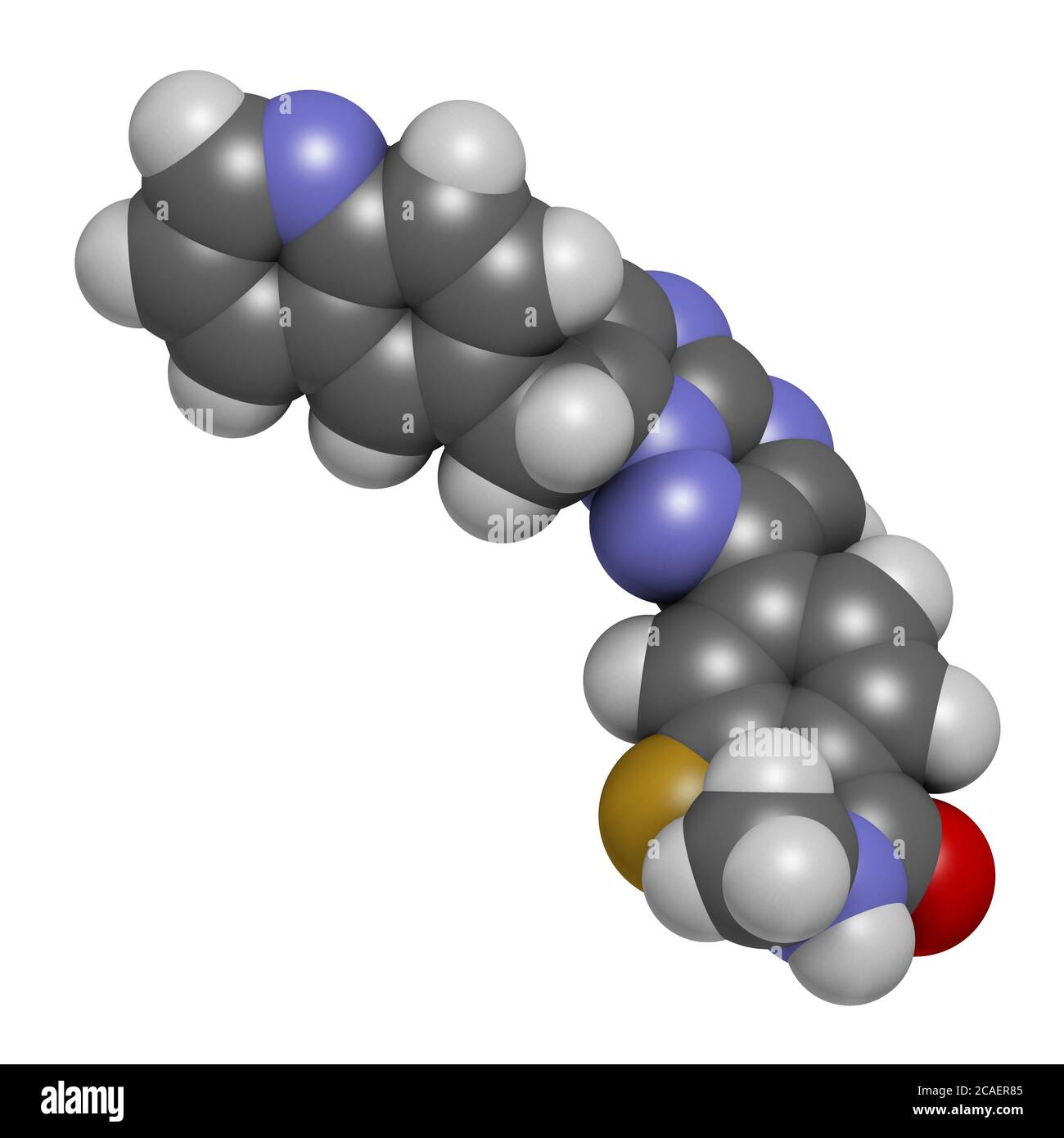
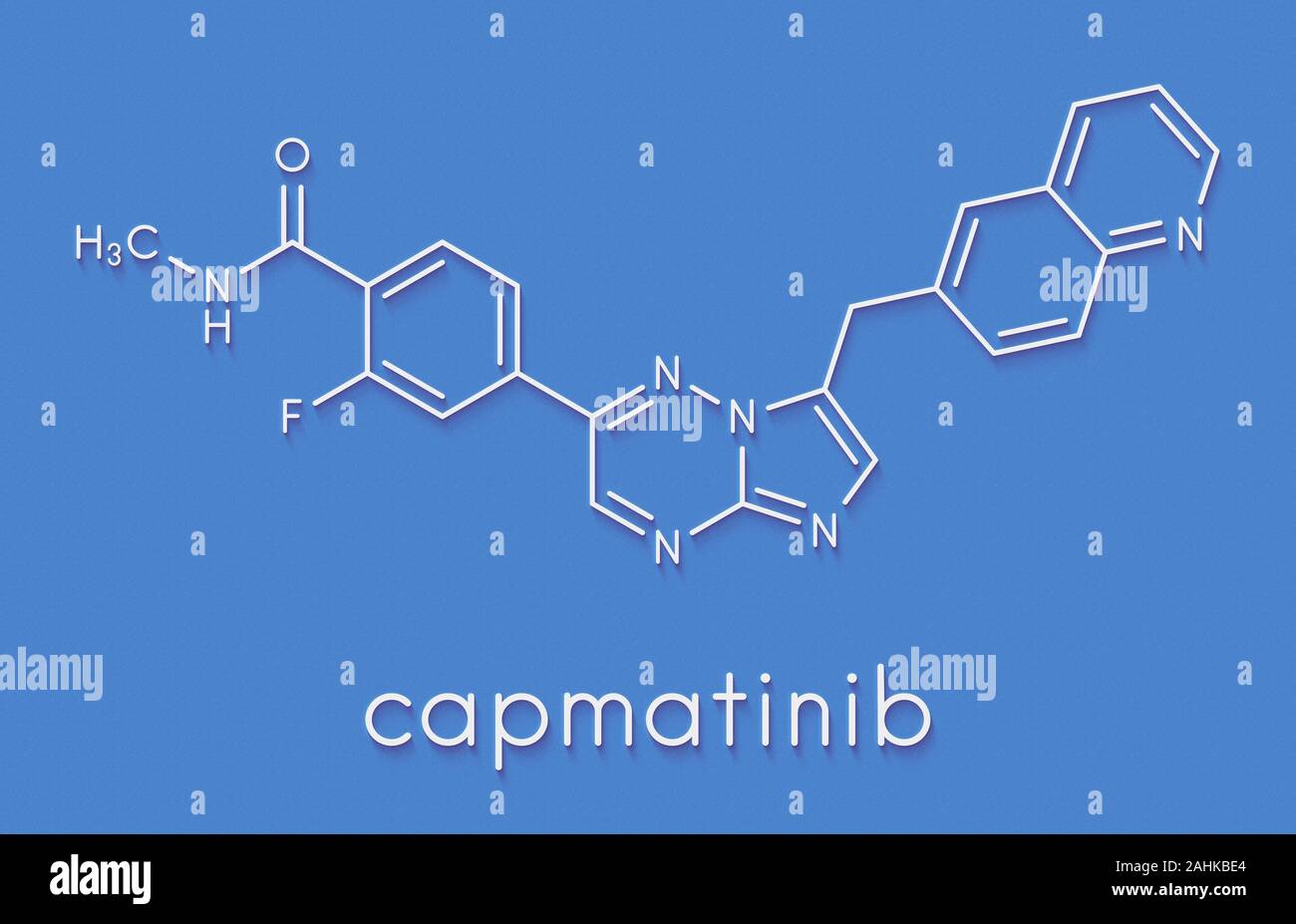
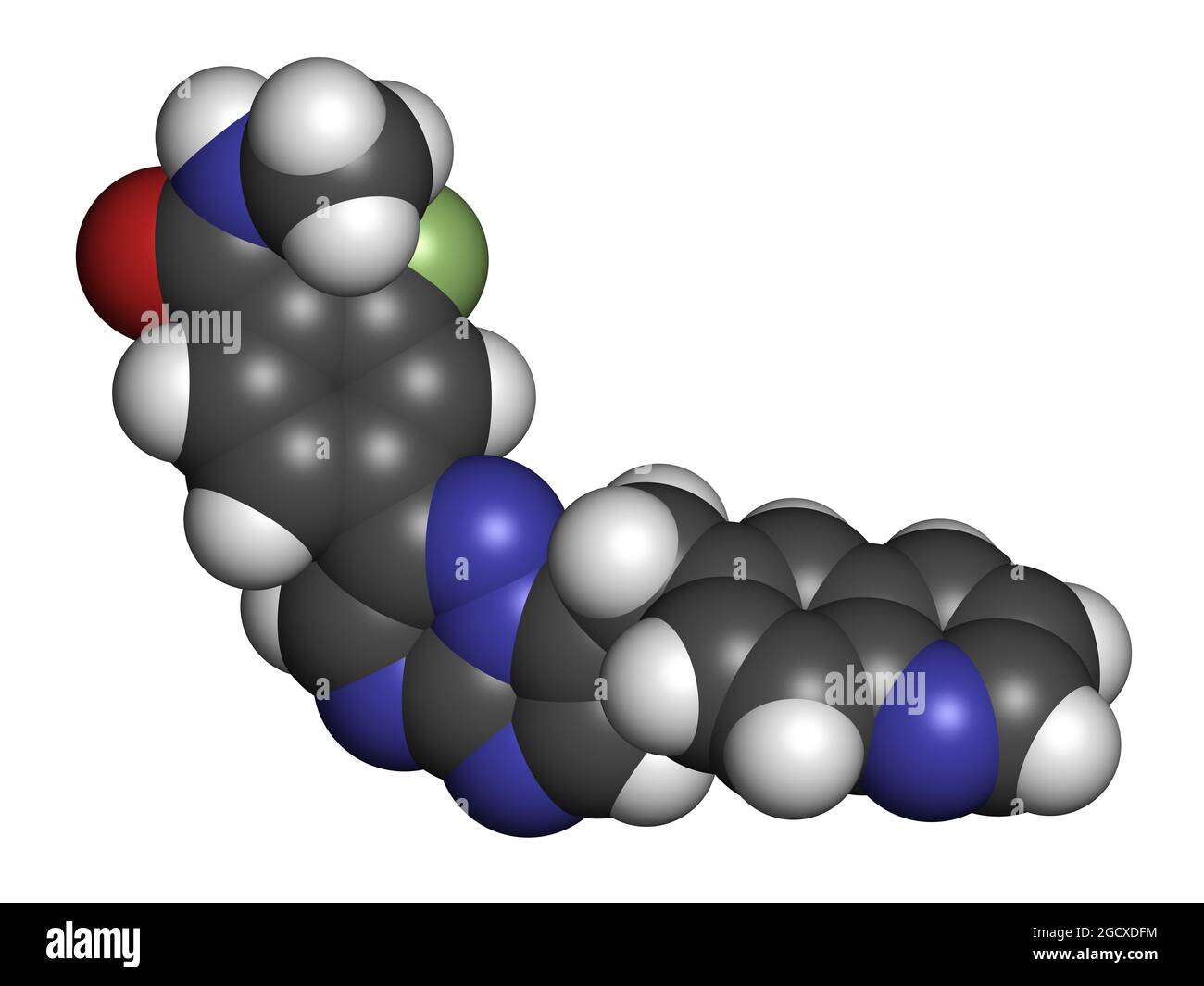
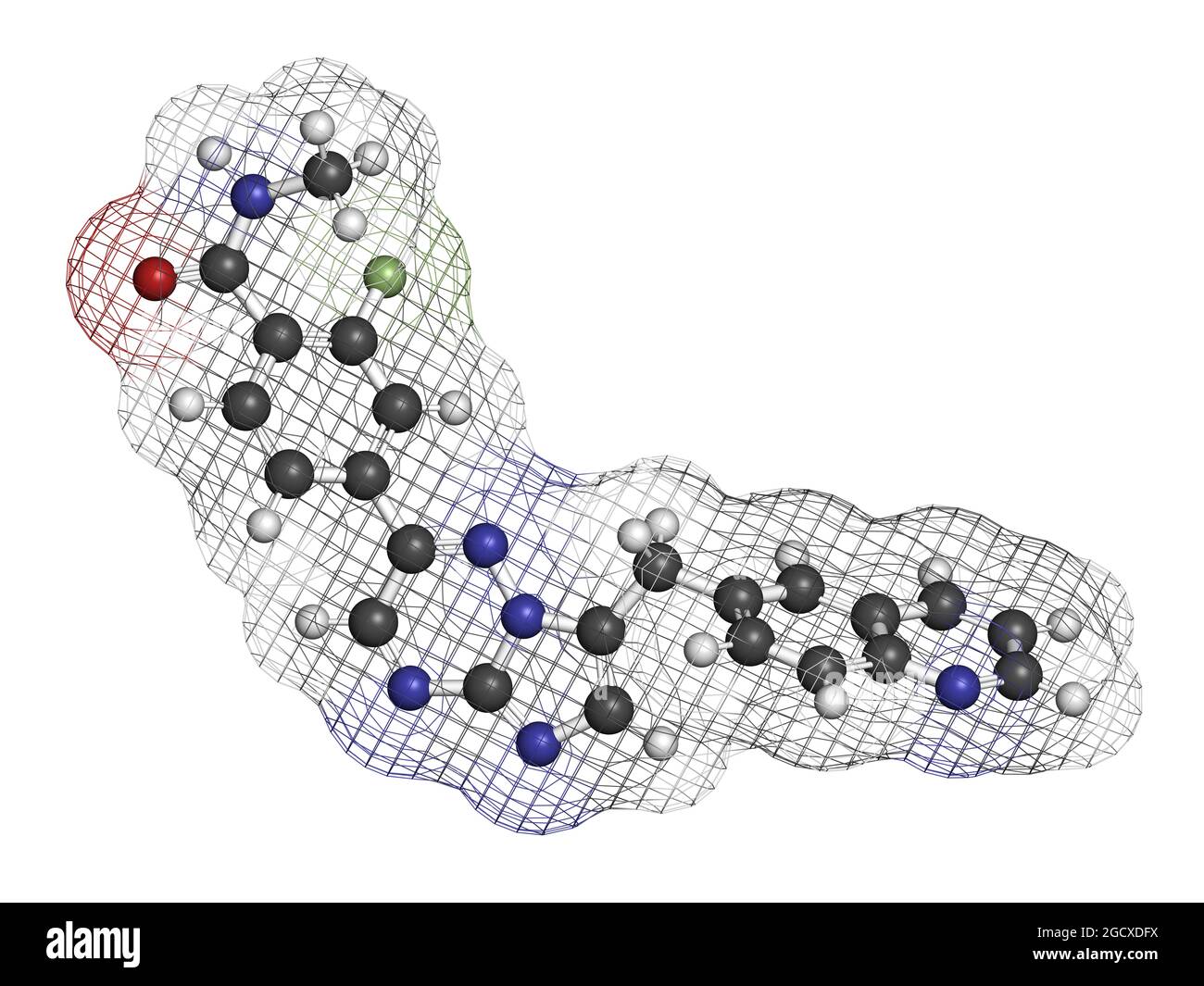
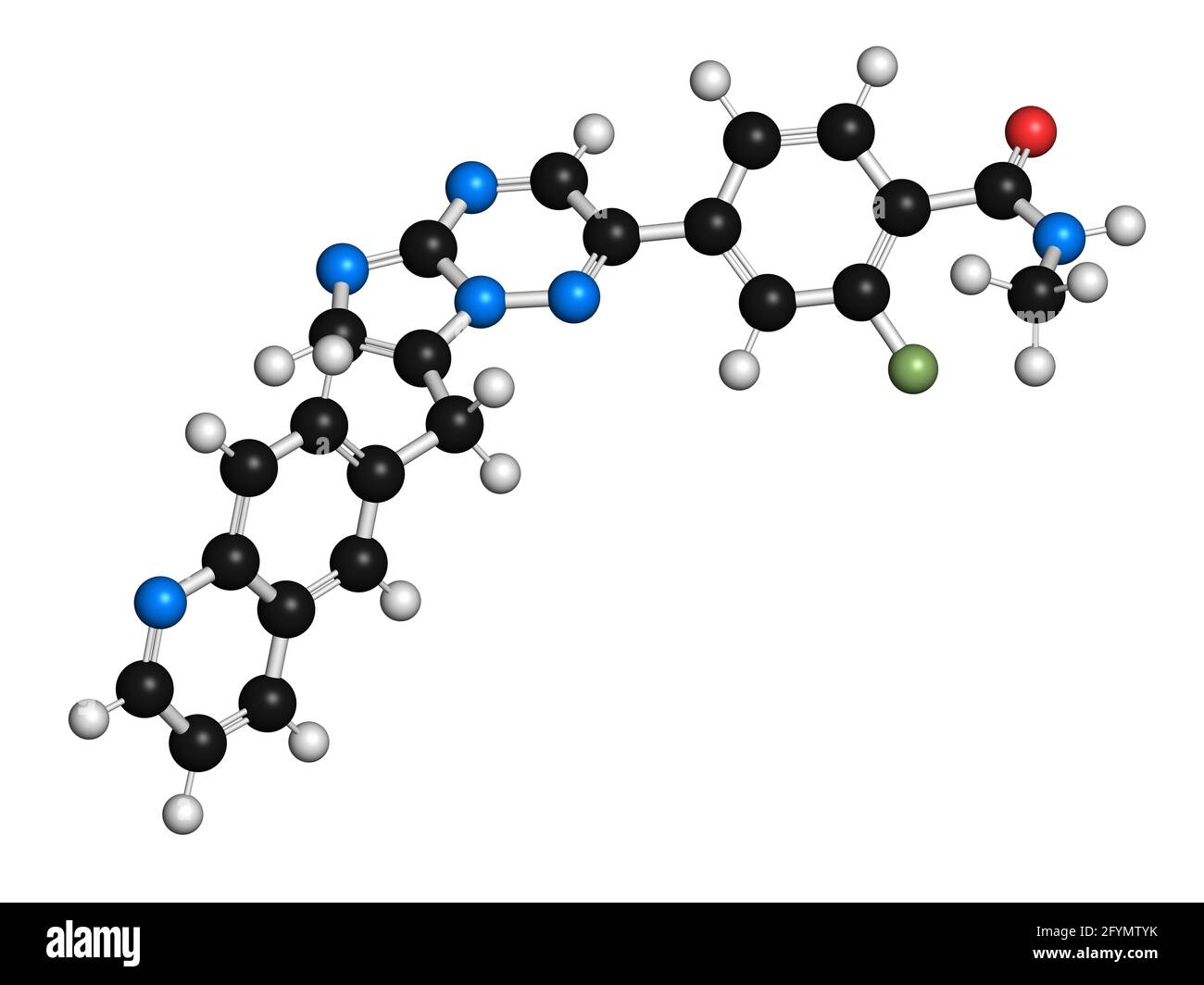
RF2CAER85–Capmatinib cancer drug molecule (c-met inhibitor). 3D rendering. Atoms are represented as spheres with conventional color coding: hydrogen (white), ca
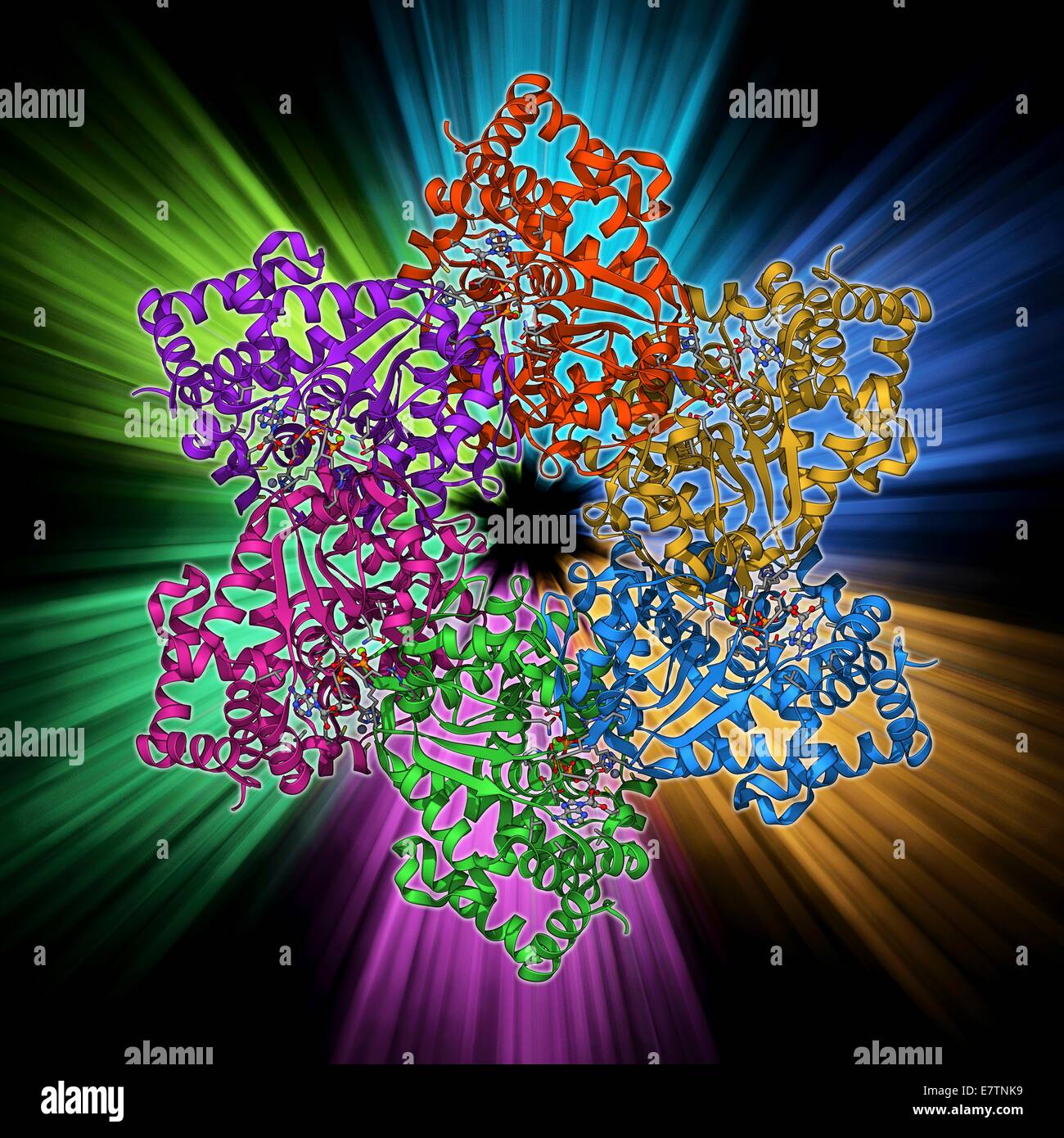
RFE7TNK9–DNA helicase. Molecular model of a helicase molecule from the SV40 virus. Helicases are enzymes that separate the two strands of the DNA double helix, by breaking the hydrogen bonds between nucleotide bases. Separation of the strands is needed for replica
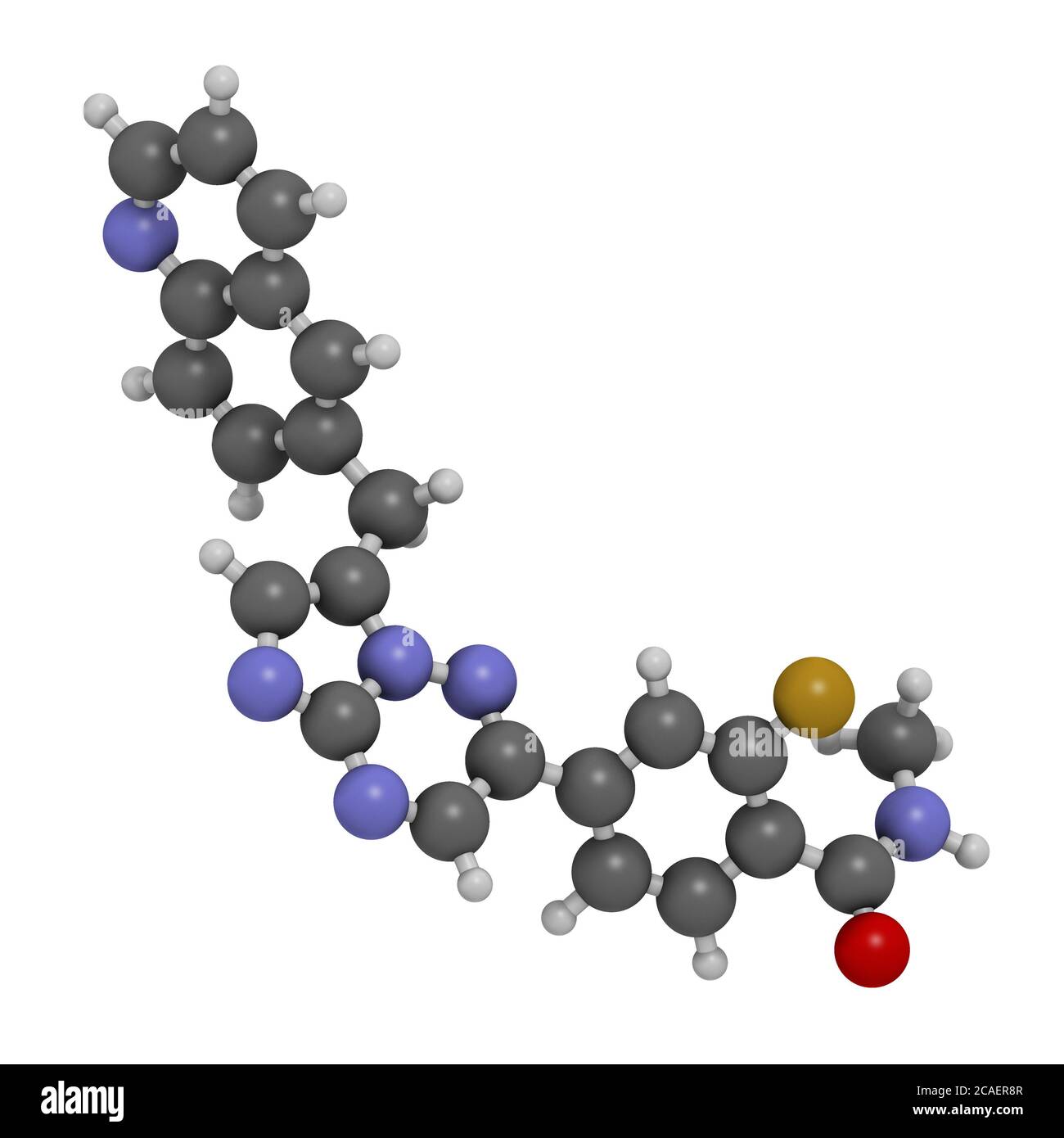
RF2CAER8R–Capmatinib cancer drug molecule (c-met inhibitor). 3D rendering. Atoms are represented as spheres with conventional color coding: hydrogen (white), ca
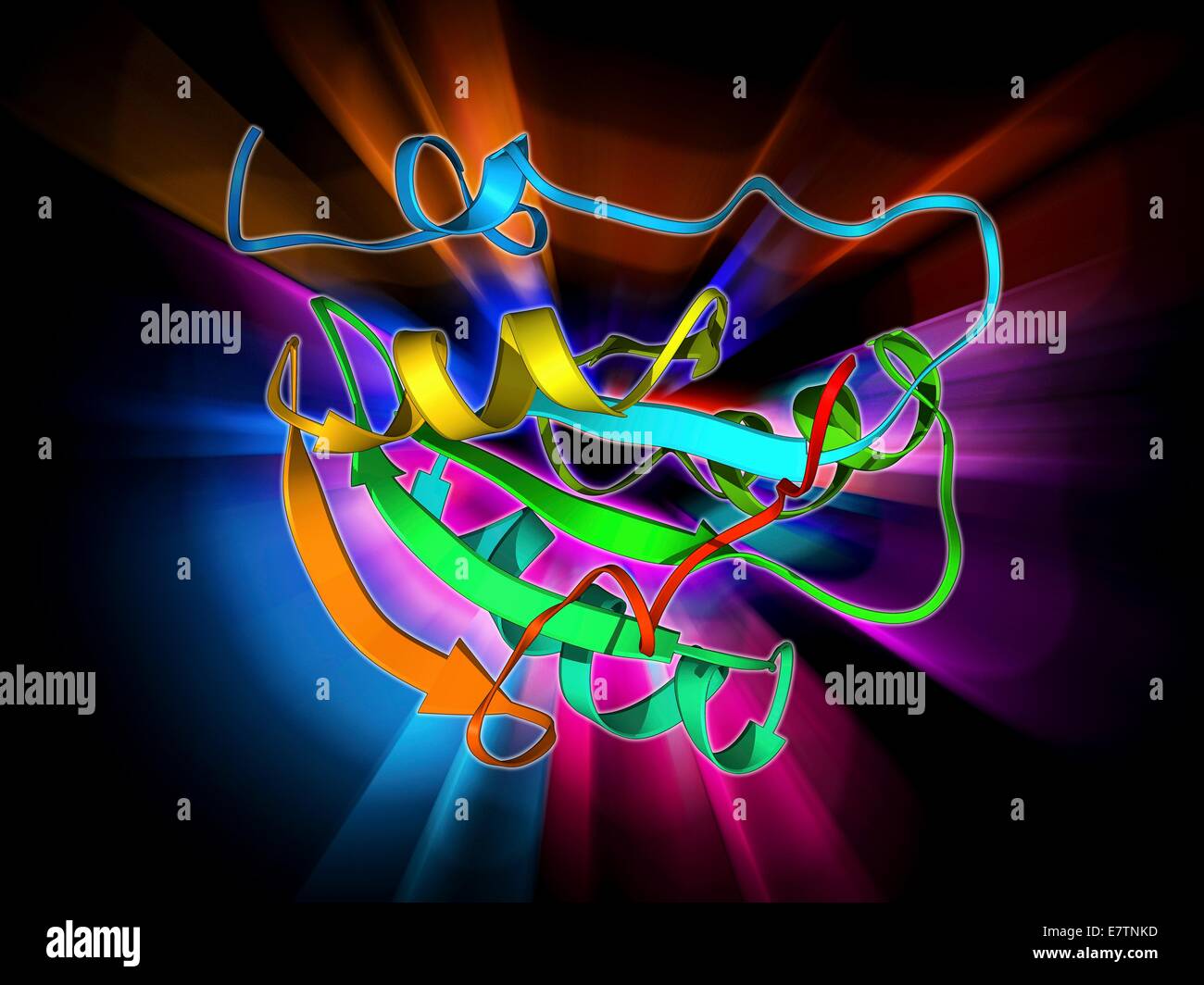
RFE7TNKD–Simian virus (SV40) large T antigen, molecular model. This antigen is from the simian vacuolating virus 40 (SV40). Large T antigens play a role in regulating the viral life cycle of the polyomaviridae viruses, such as SV40. SV40 is found in monkeys such a
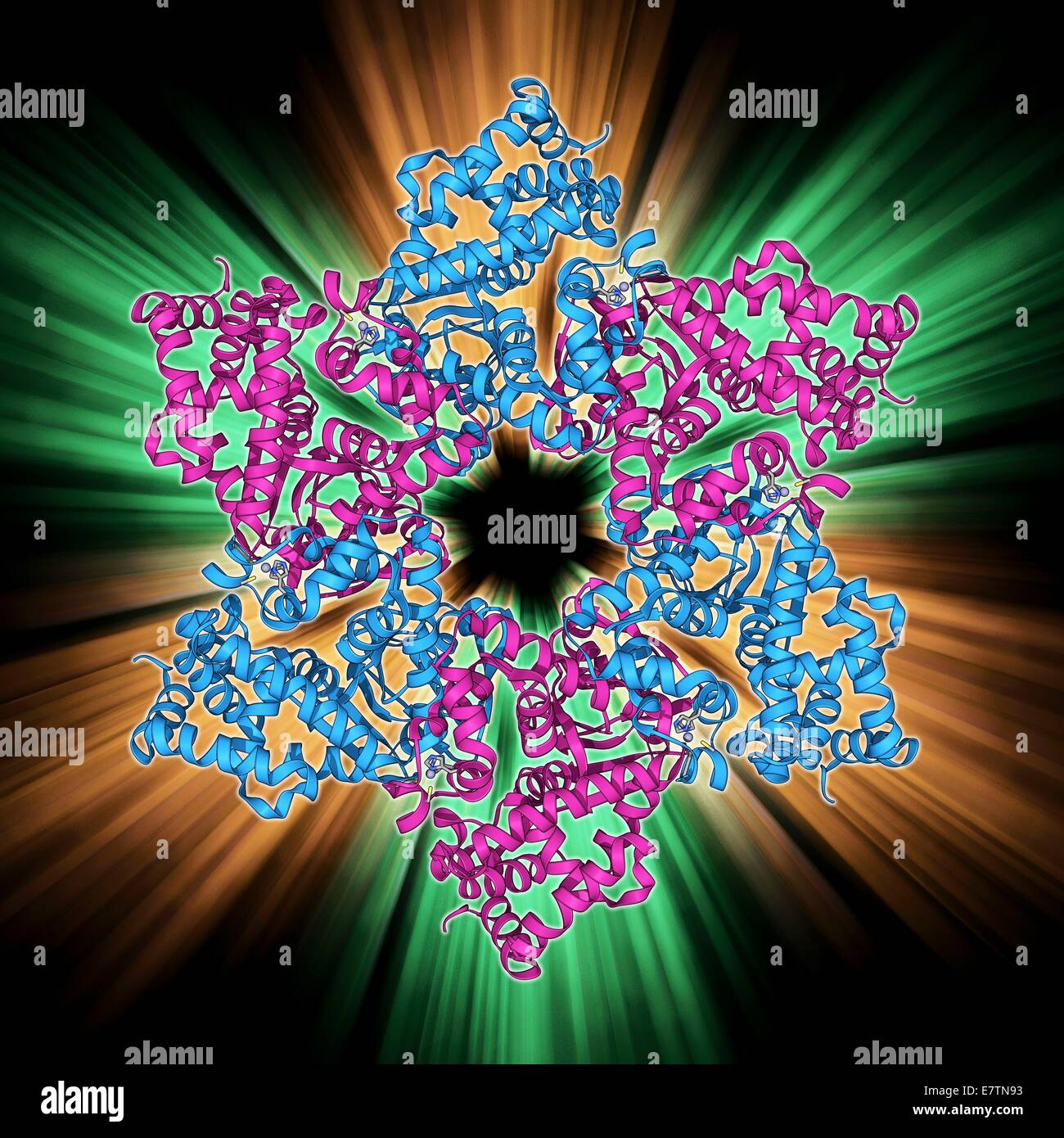

RFE7TN93–DNA helicase. Molecular model of a helicase molecule from the SV40 virus. Helicases are enzymes that separate the two strands of the DNA double helix, by breaking the hydrogen bonds between nucleotide bases. Separation of the strands is needed for replica
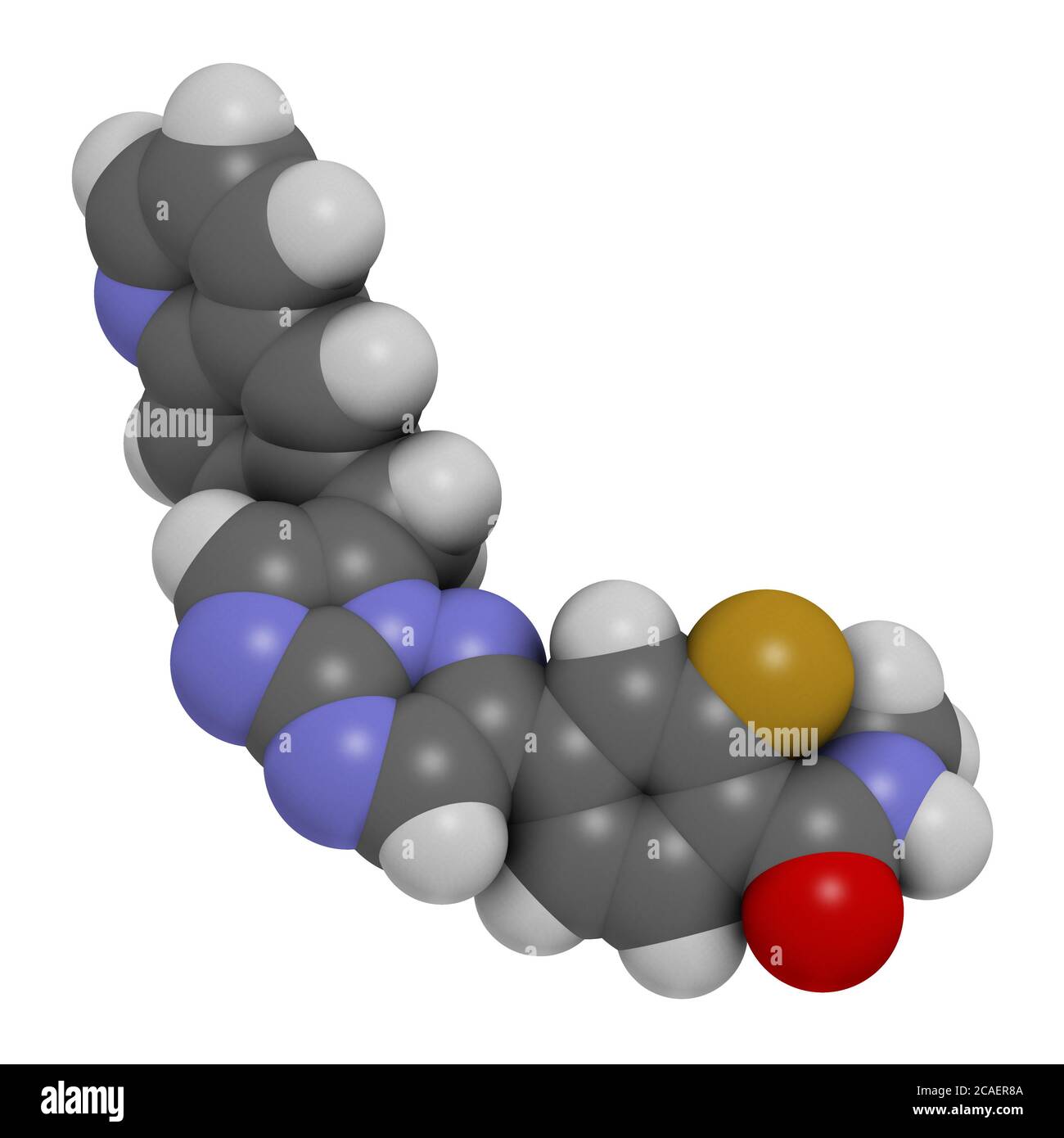
RF2CAER8A–Capmatinib cancer drug molecule (c-met inhibitor). 3D rendering. Atoms are represented as spheres with conventional color coding: hydrogen (white), ca
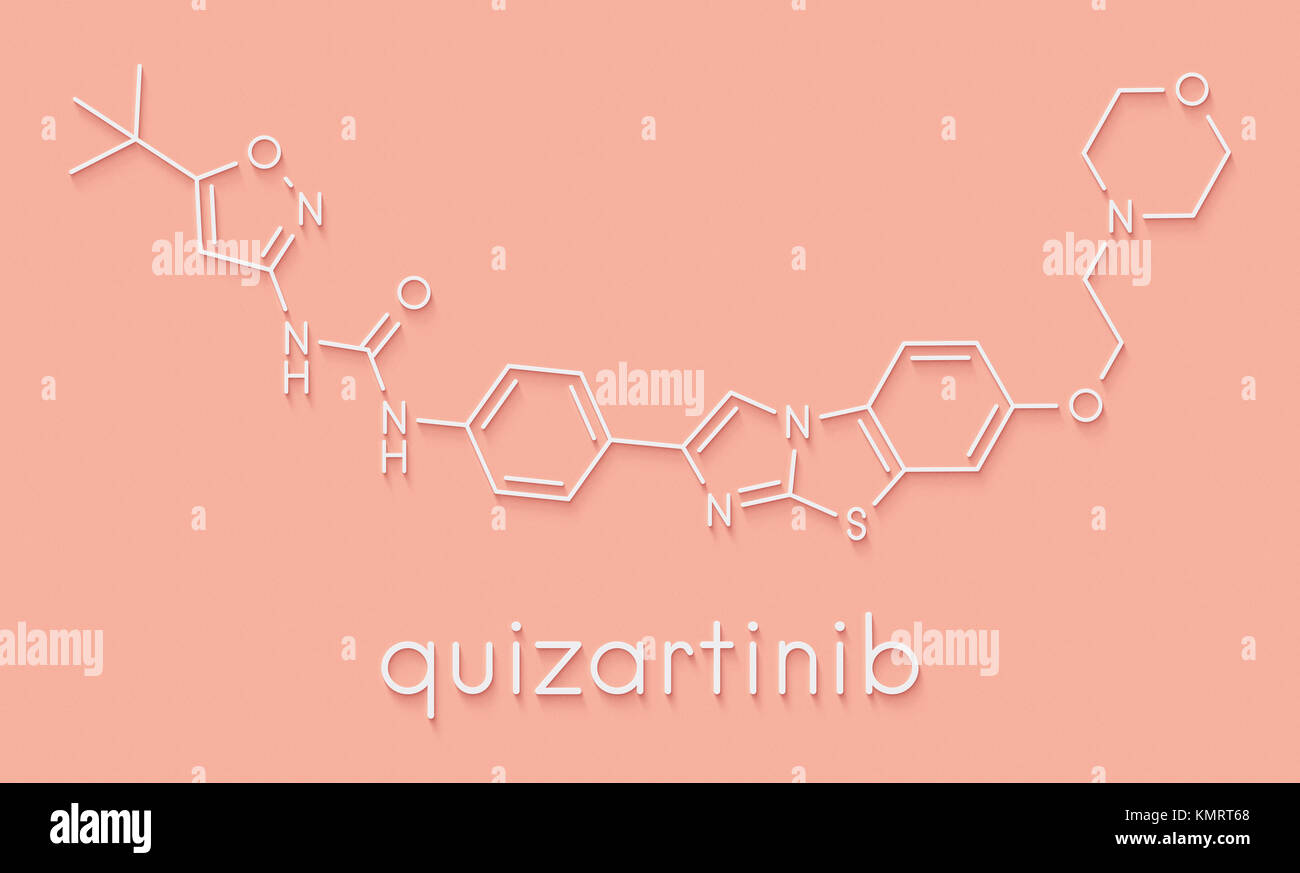
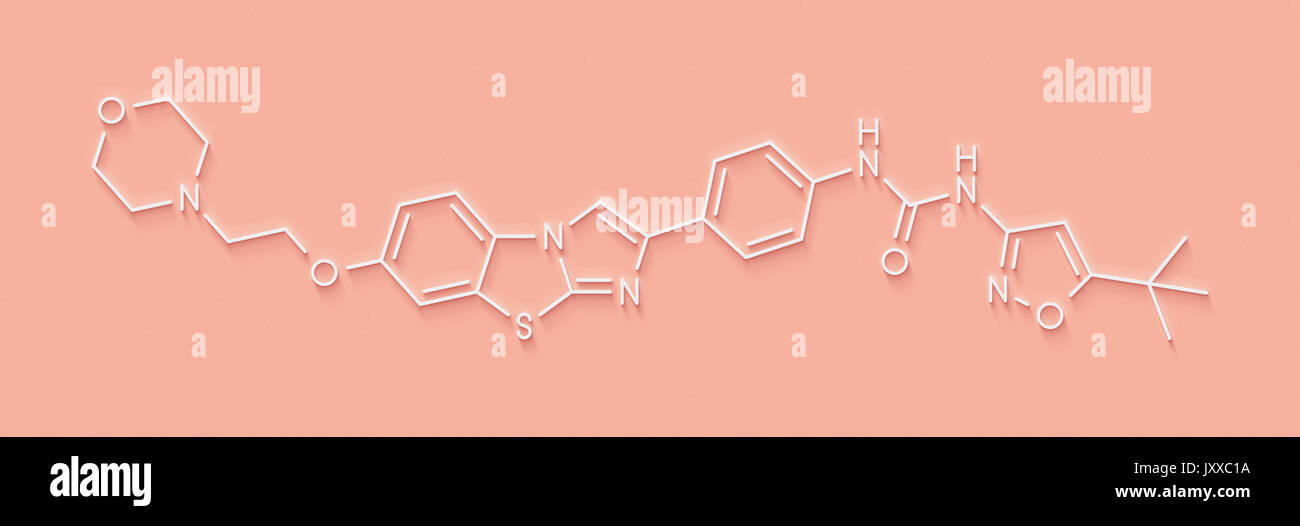
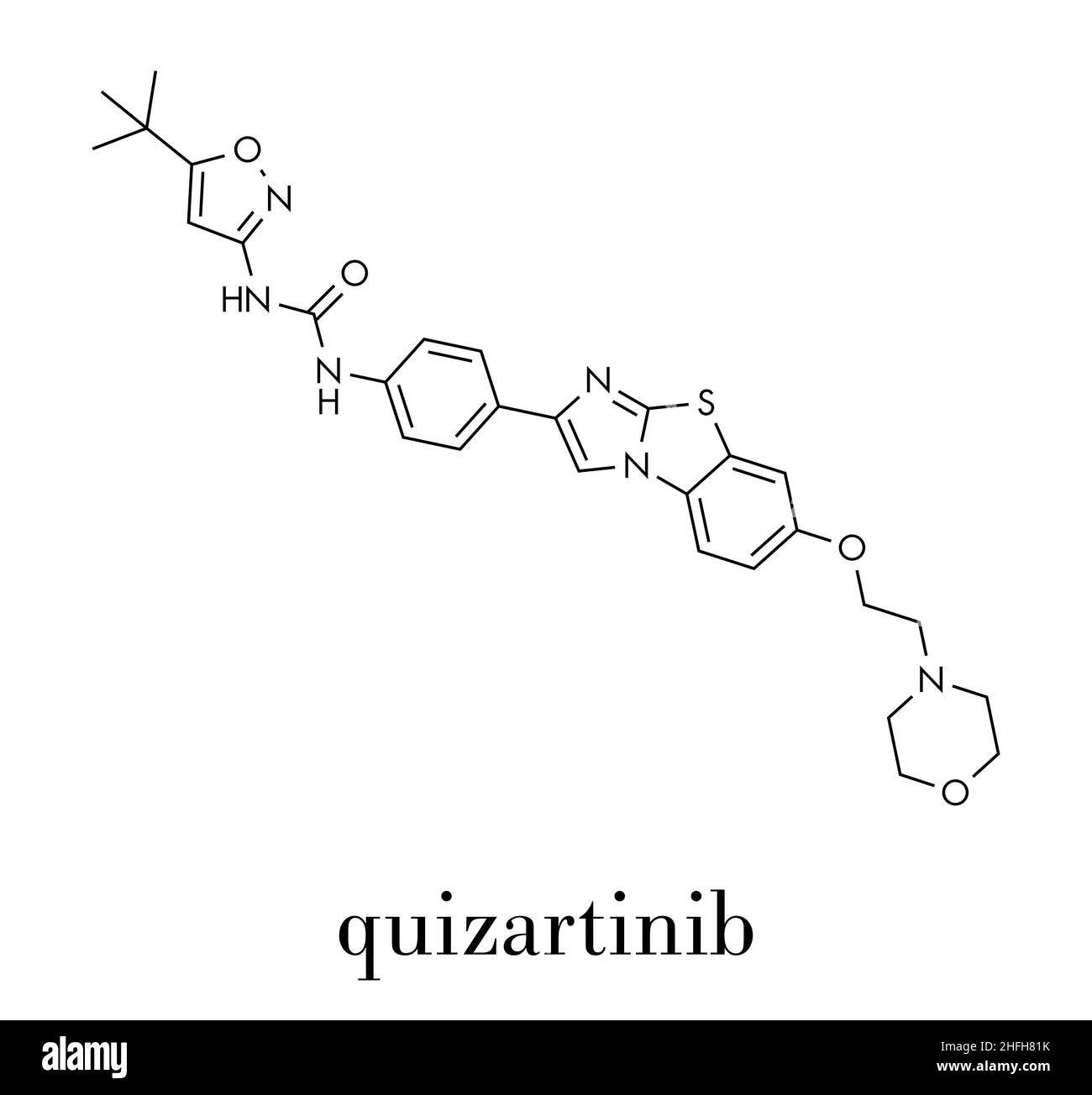
RFKMRT68–Quizartinib investigational acute myeloid leukemia (AML) drug, chemical structure Skeletal formula.
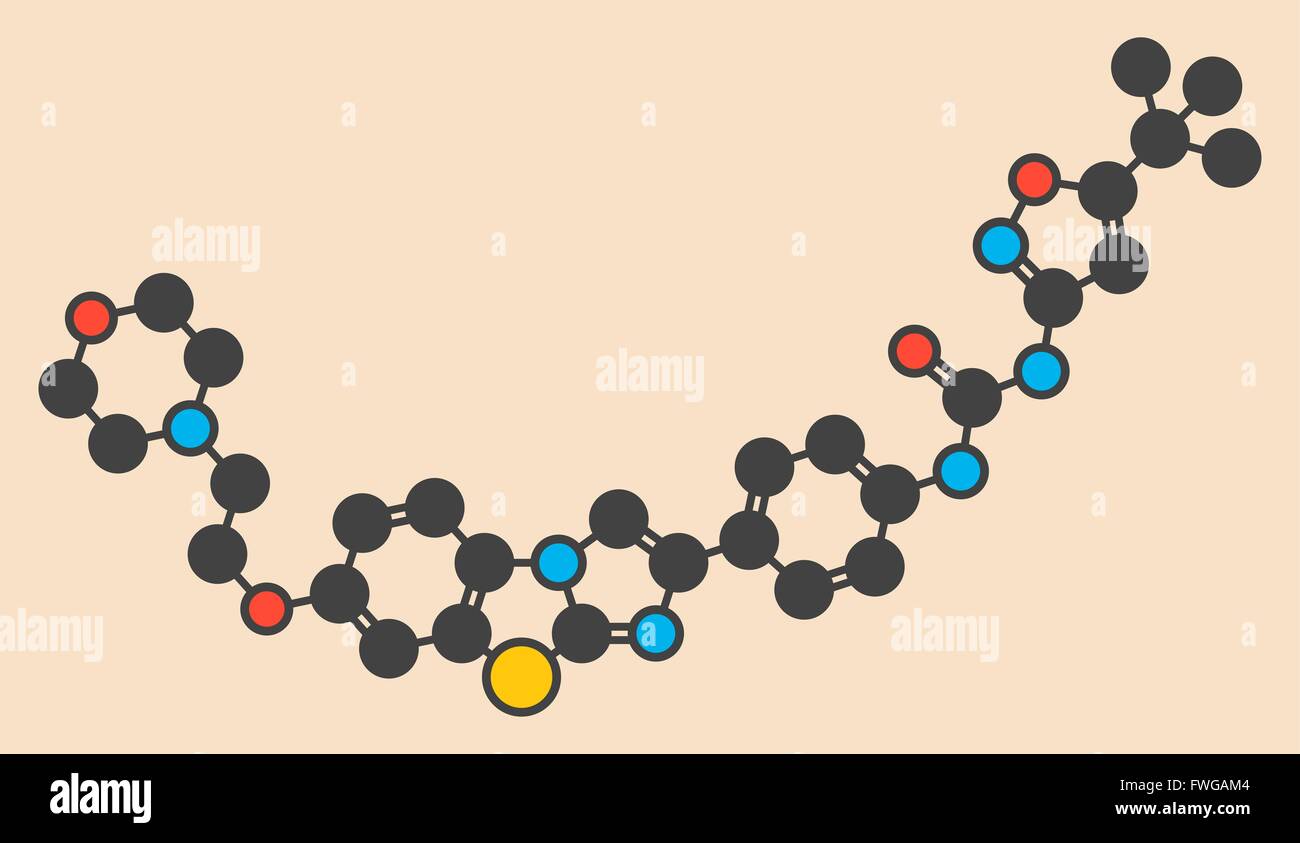
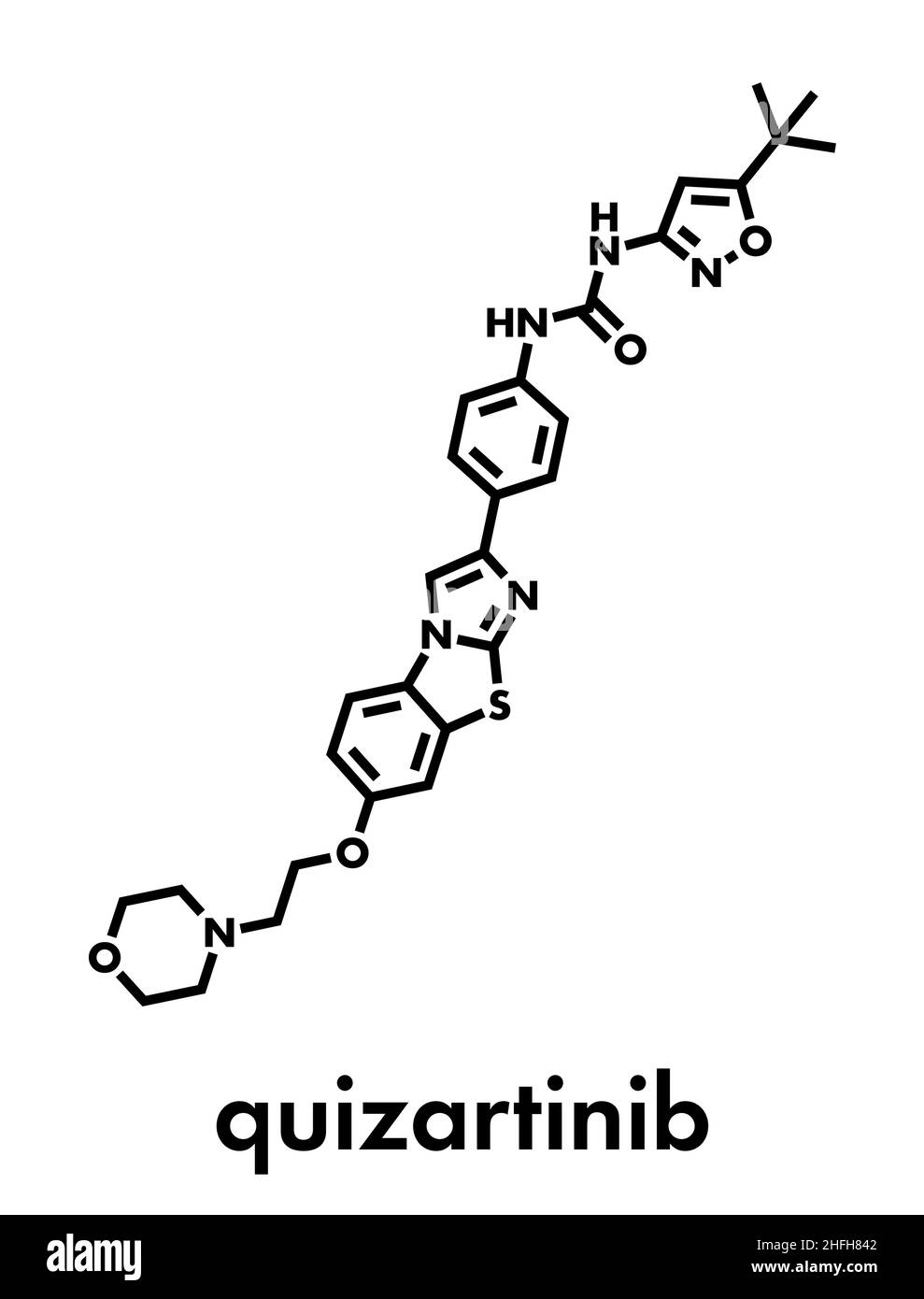
RFFWGAM4–Quizartinib investigational acute myeloid leukaemia (AML) drug chemical structure Stylized skeletal formula (chemical
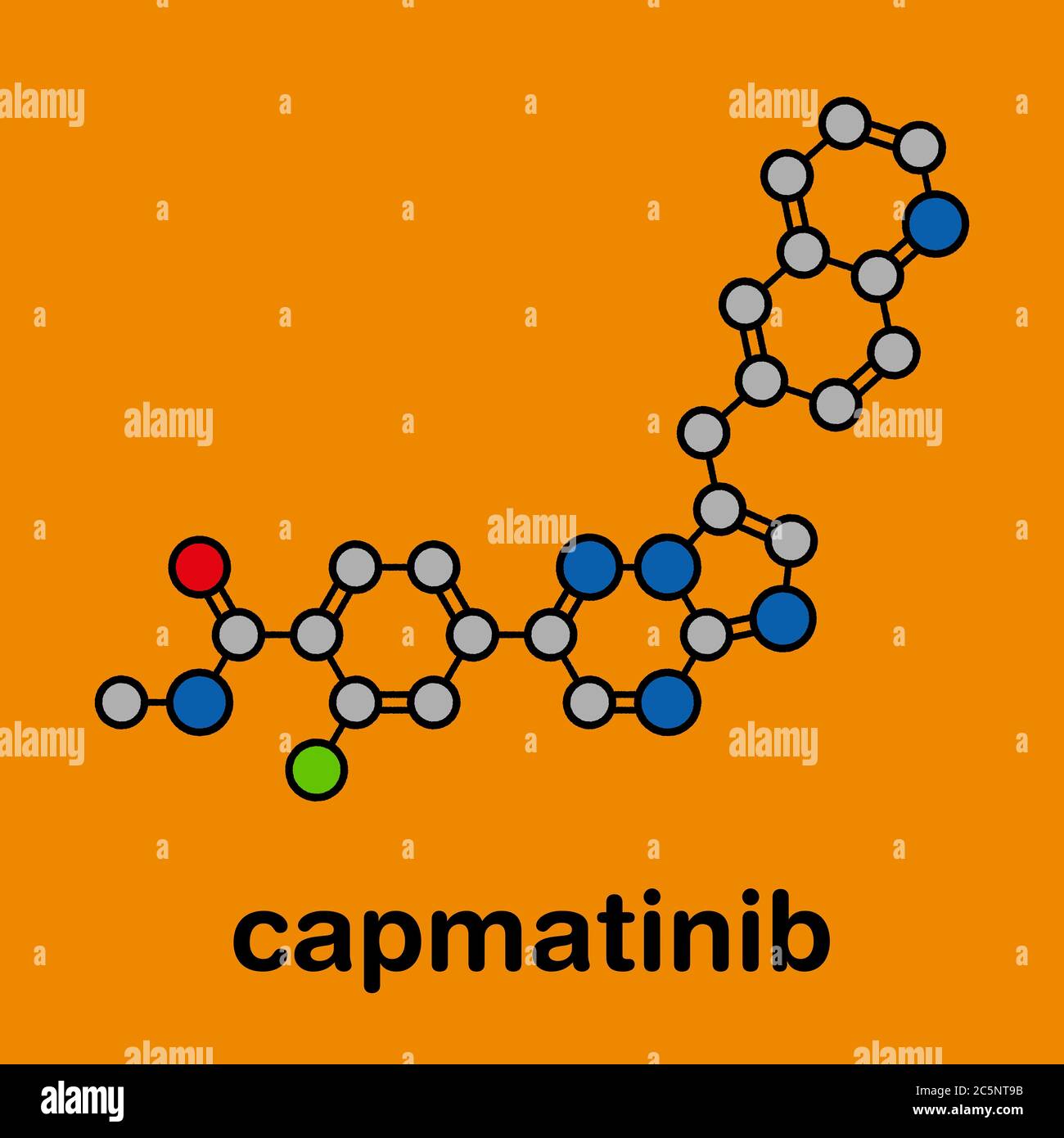

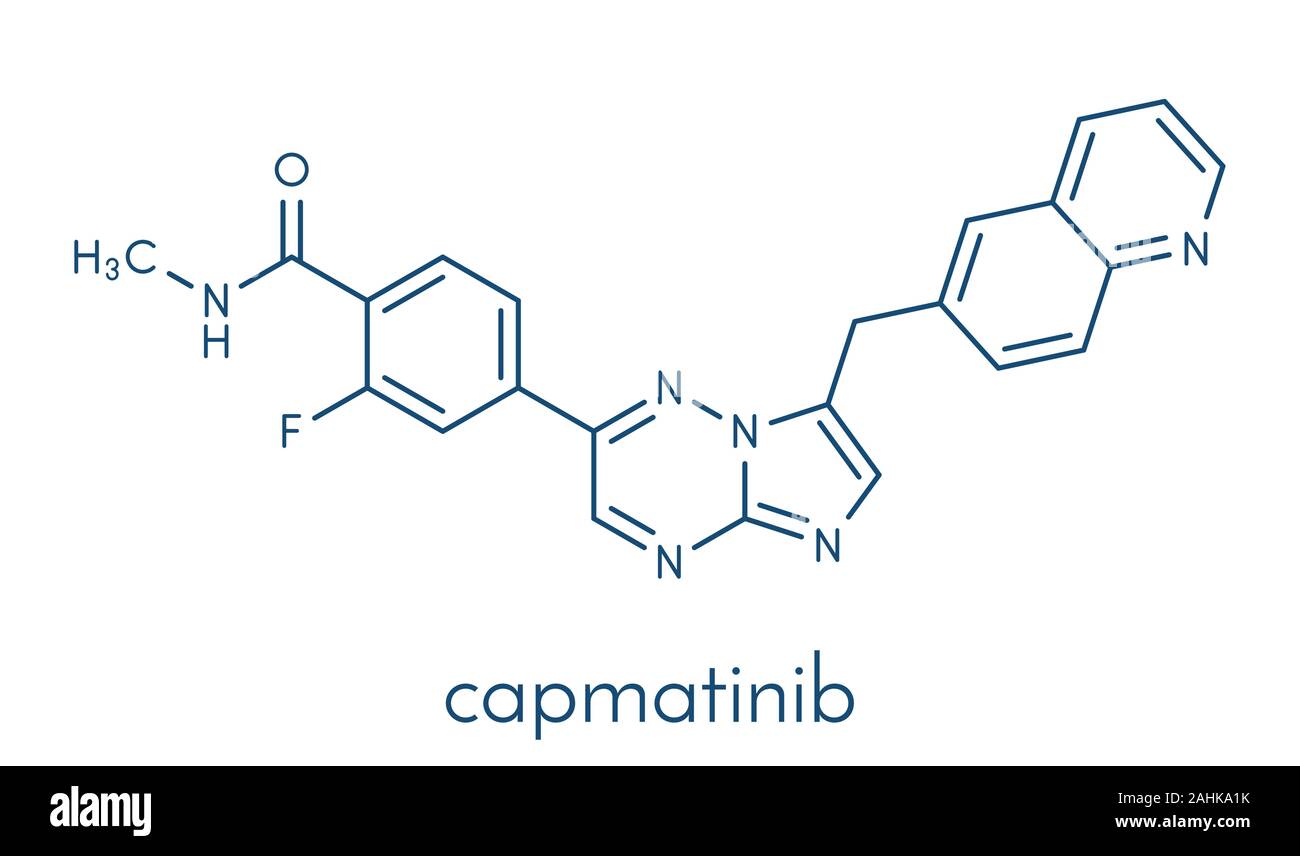
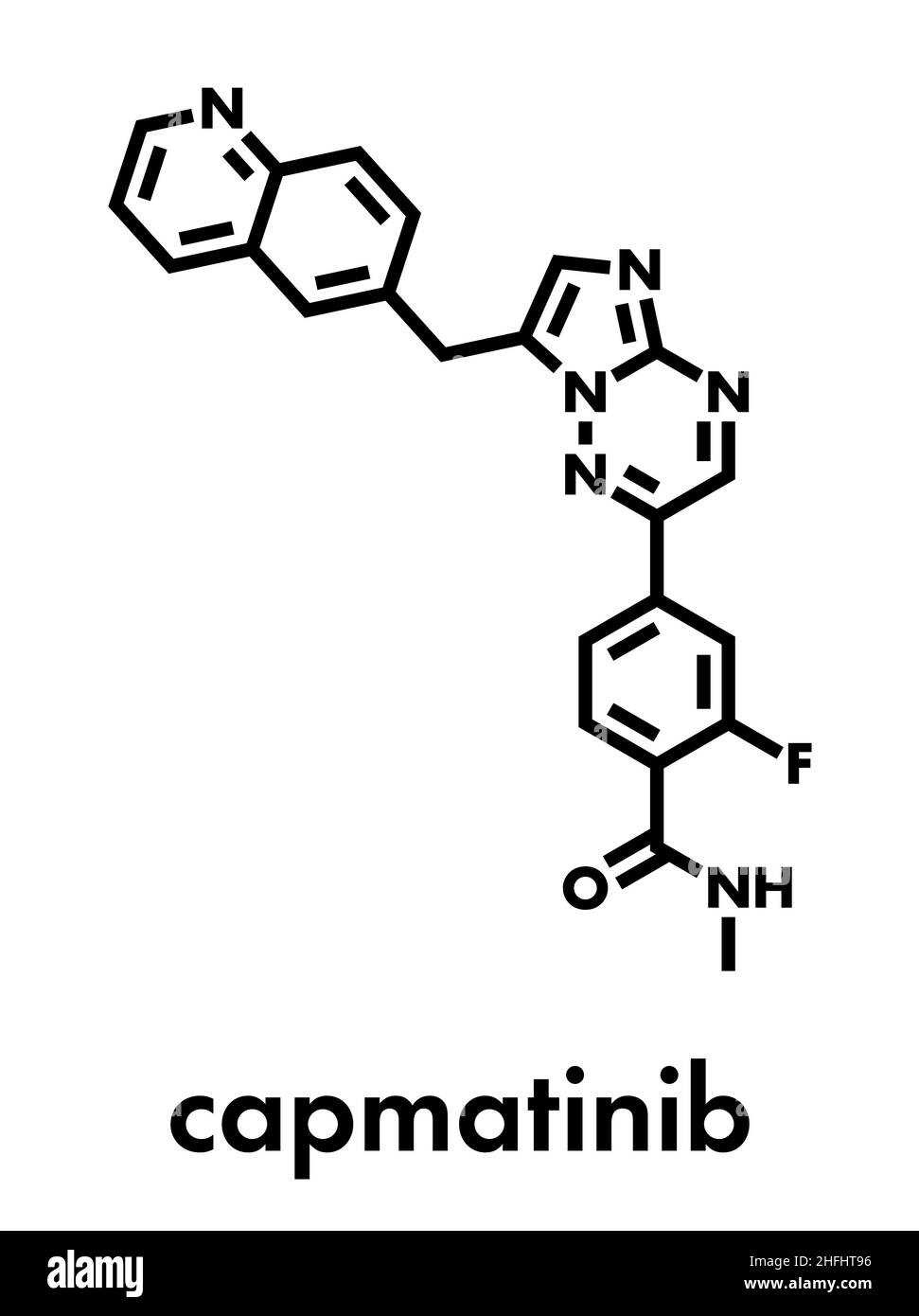
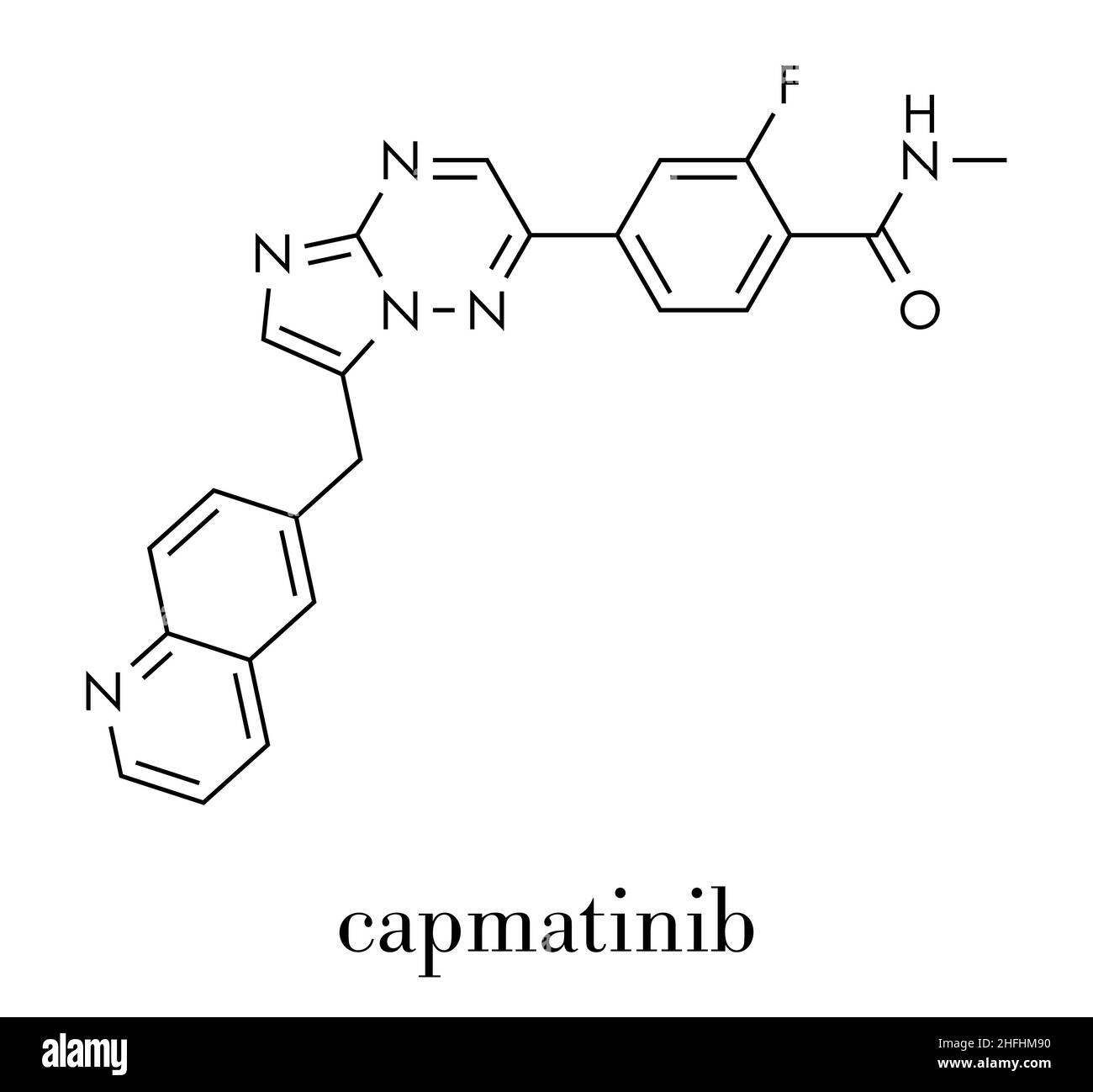
RF2C5NT9B–Capmatinib cancer drug molecule (c-met inhibitor). Stylized skeletal formula (chemical structure): Atoms are shown as color-coded circles: hydrogen (hidden), carbon (grey), nitrogen (blue), oxygen (red), fluorine (cyan).
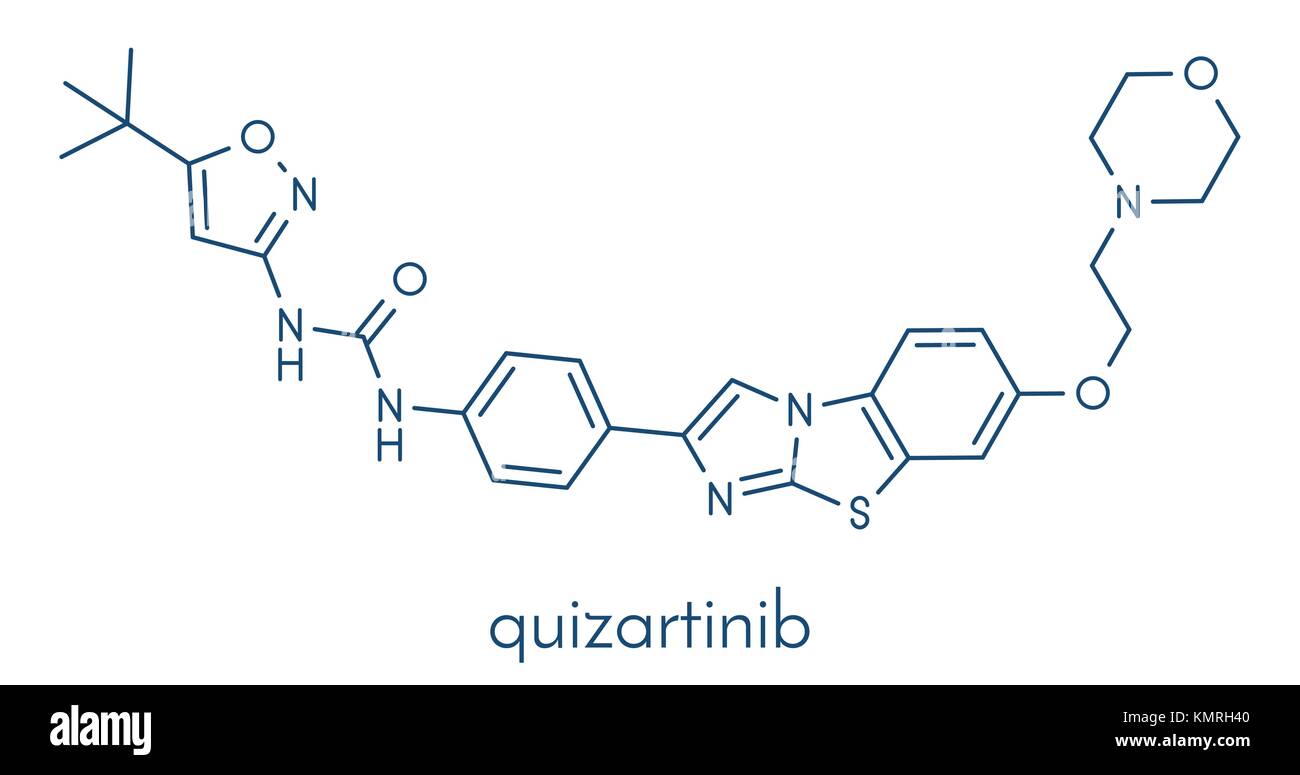
RFKMRH40–Quizartinib investigational acute myeloid leukemia (AML) drug, chemical structure Skeletal formula.
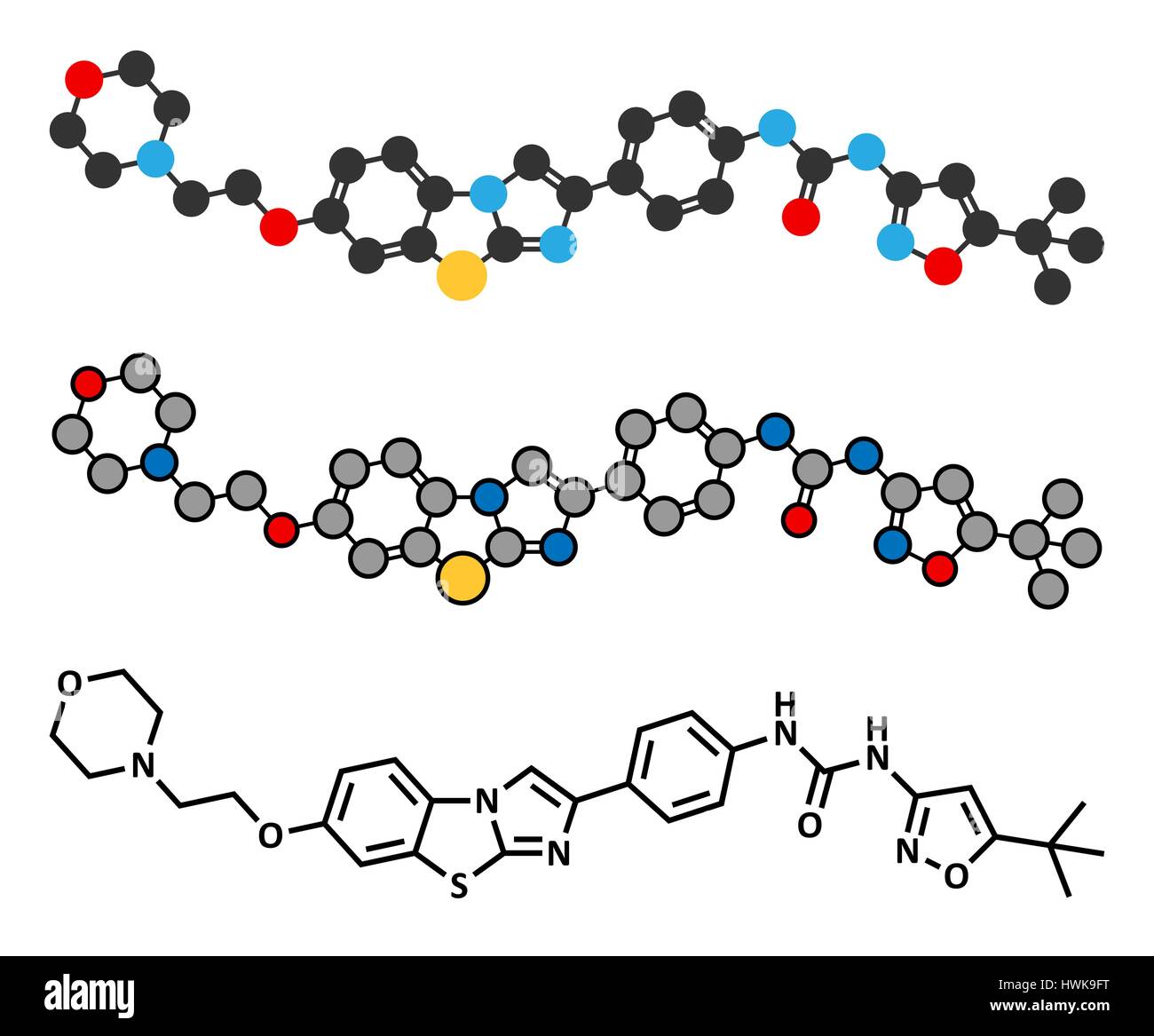
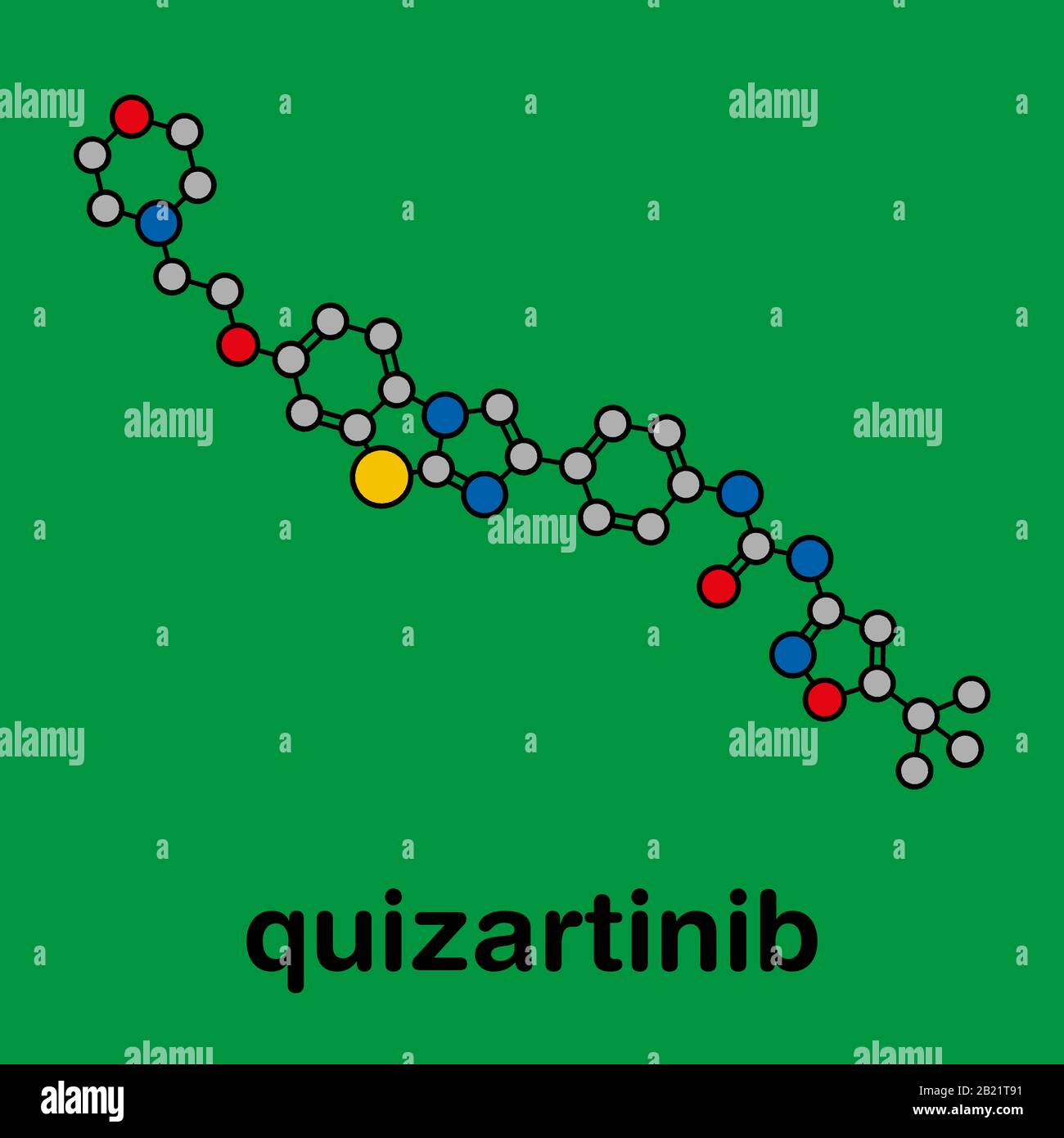
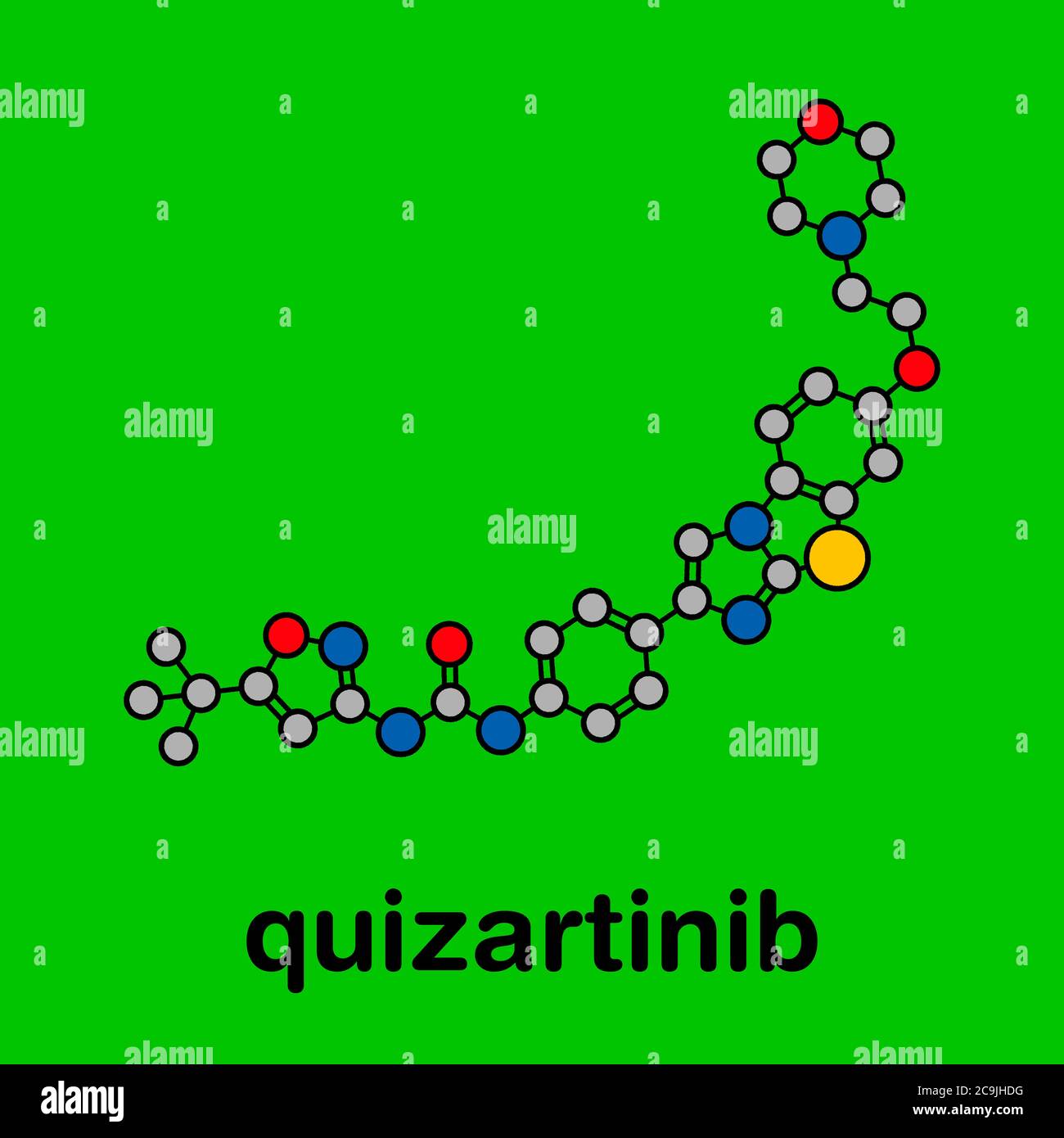
RFHWK9FT–Quizartinib cancer drug molecule (kinase inhibitor). Stylized 2D renderings and conventional skeletal formula.
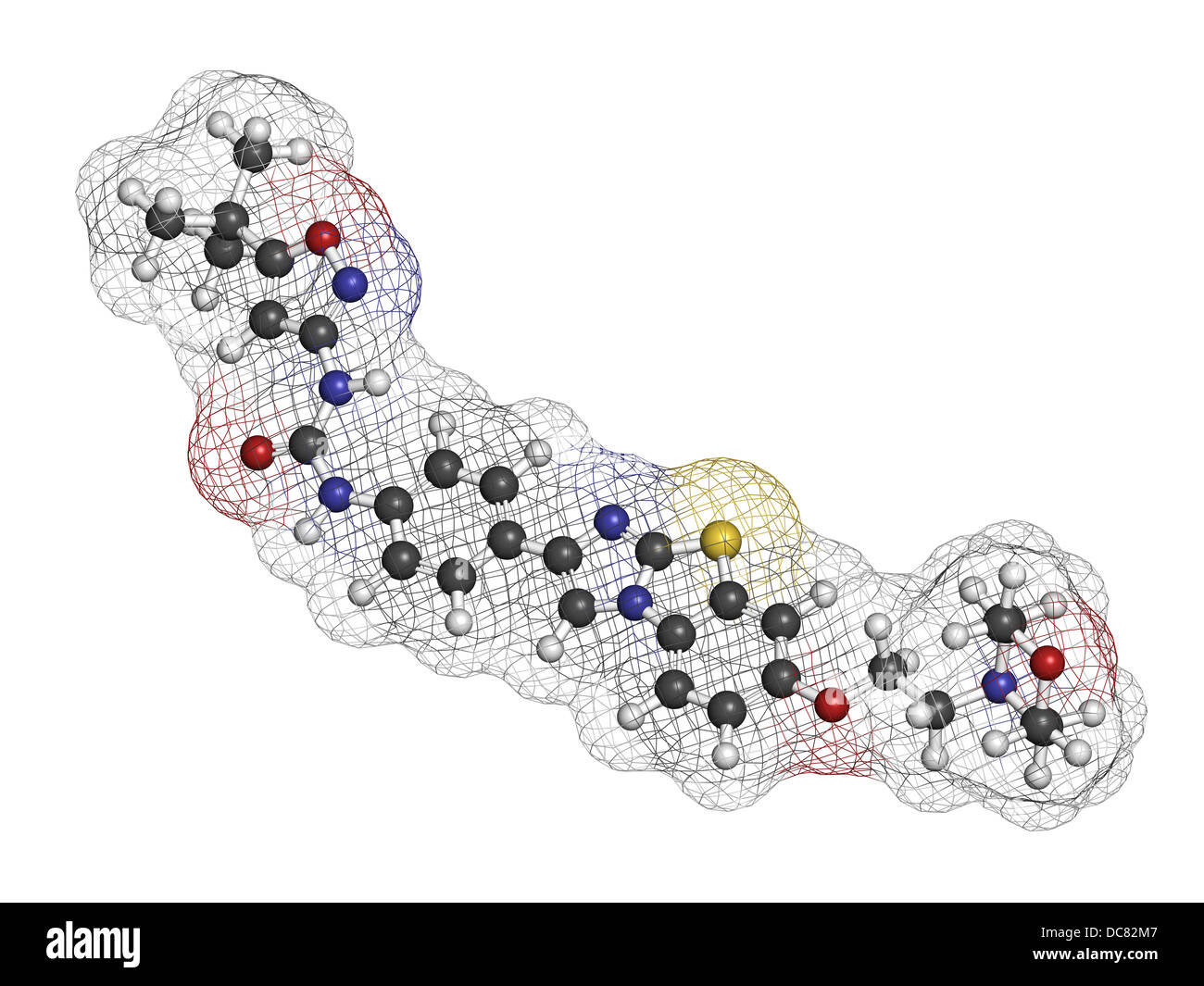
RFDC82M7–Quizartinib investigational acute myeloid leukemia (AML) drug, chemical structure Atoms are represented as spheres.
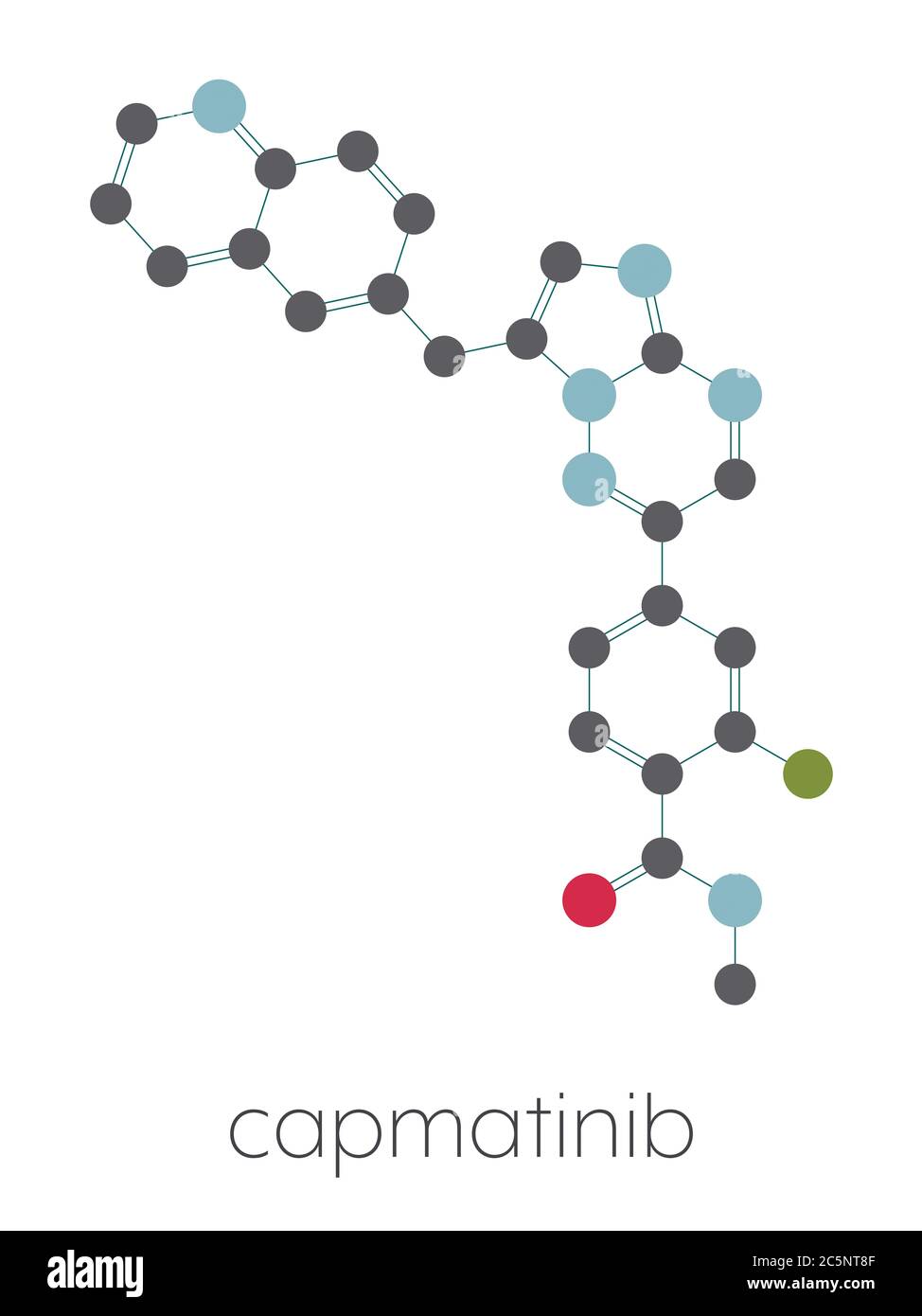
RF2C5NT8F–Capmatinib cancer drug molecule (c-met inhibitor). Stylized skeletal formula (chemical structure): Atoms are shown as color-coded circles: hydrogen (hidden), carbon (grey), nitrogen (blue), oxygen (red), fluorine (cyan).
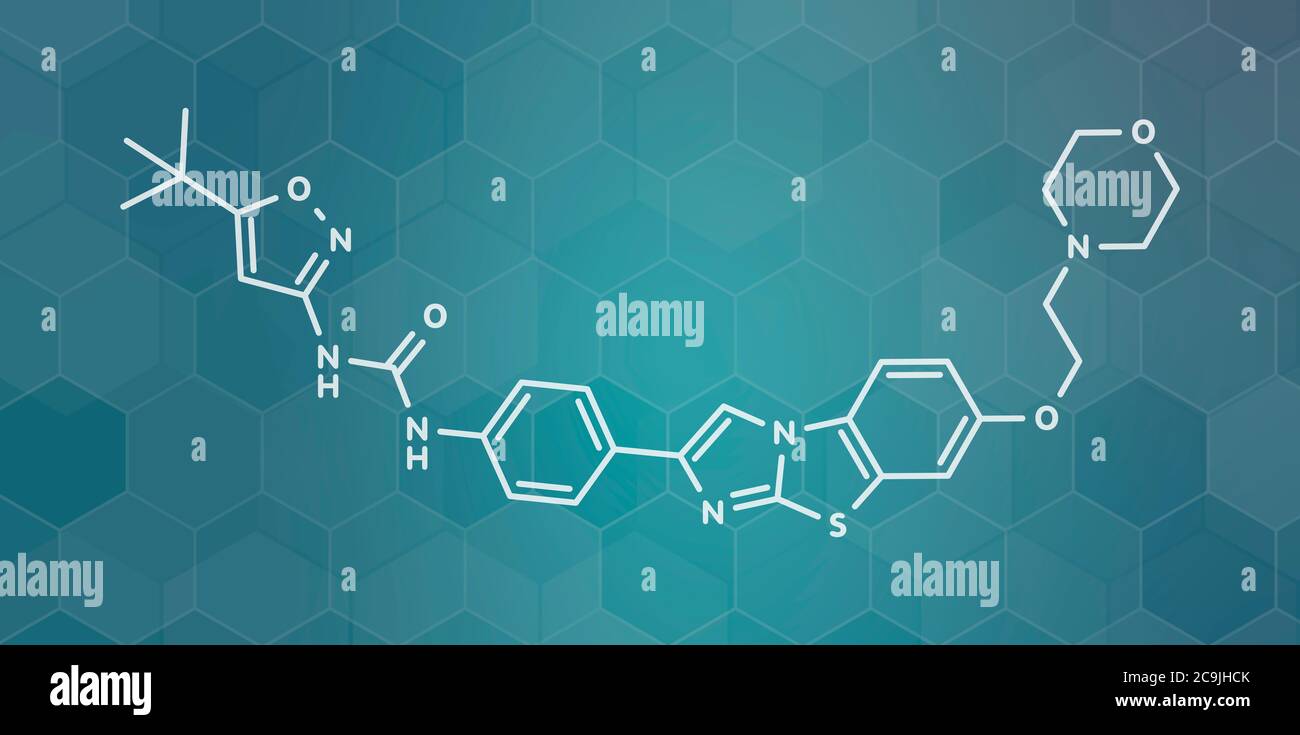
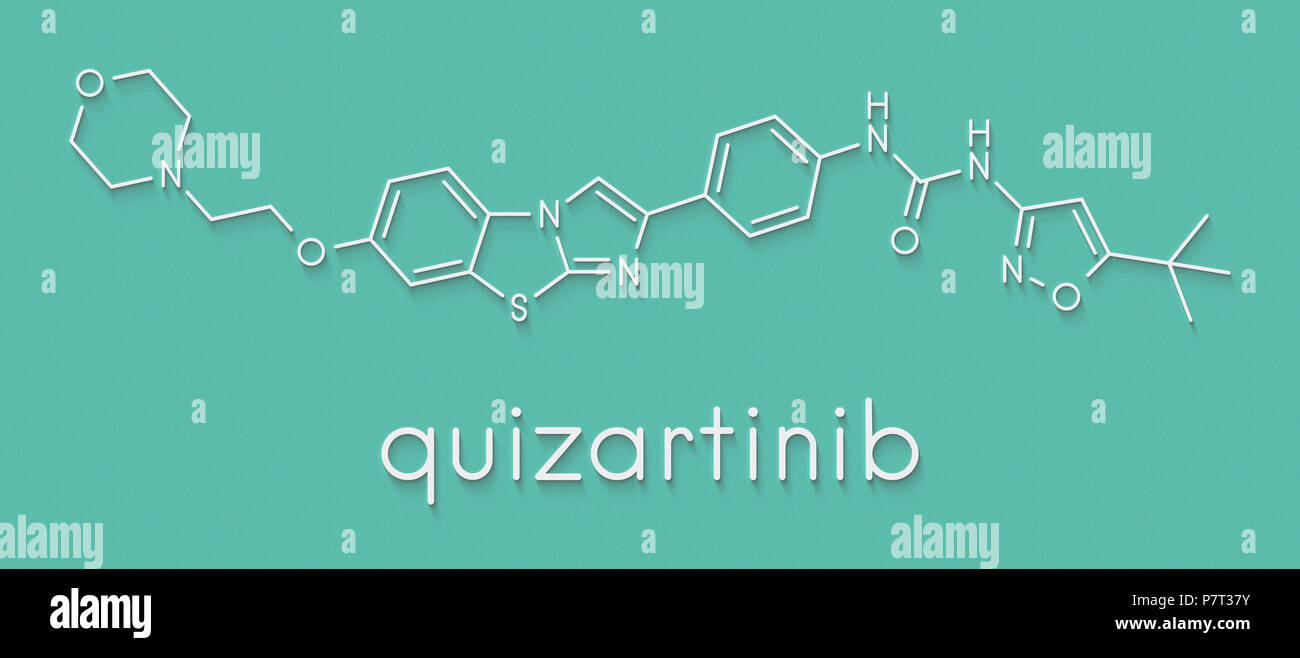
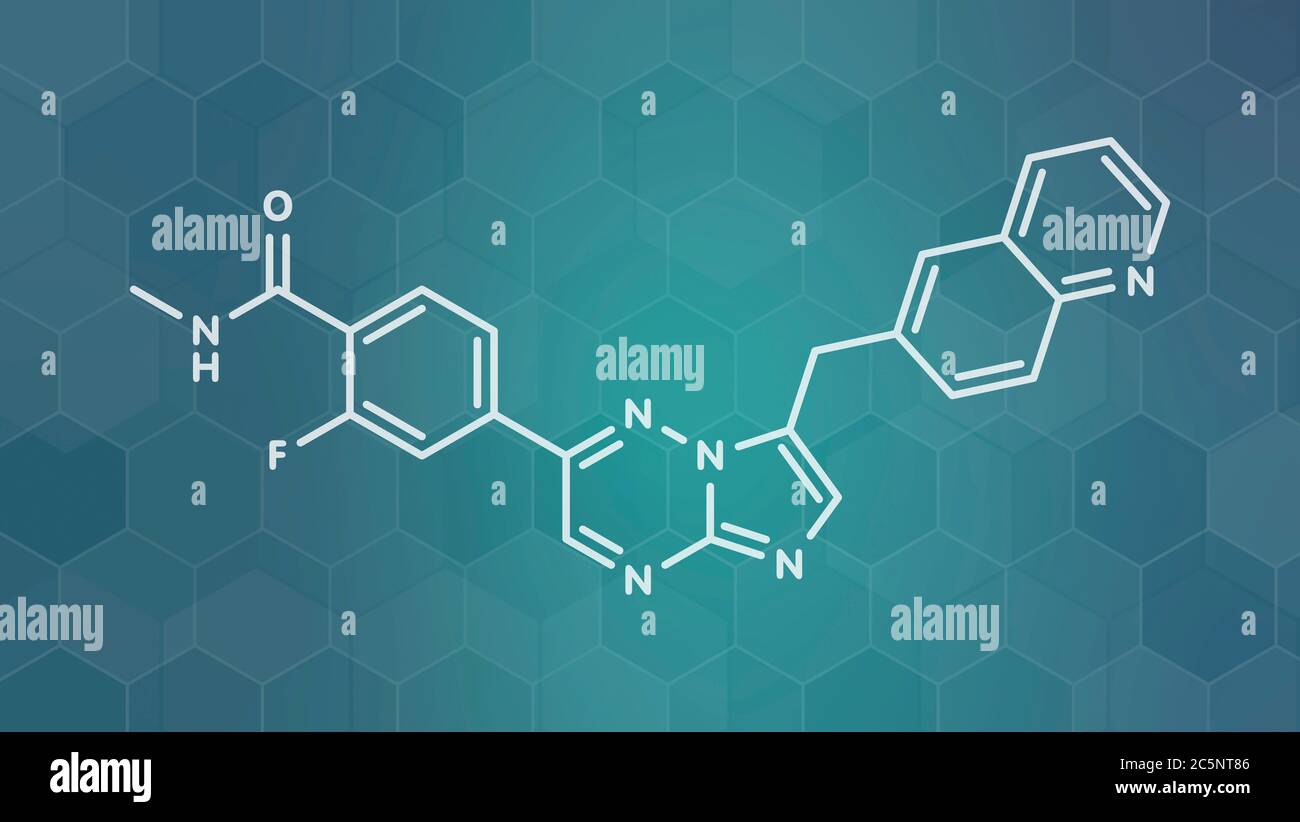
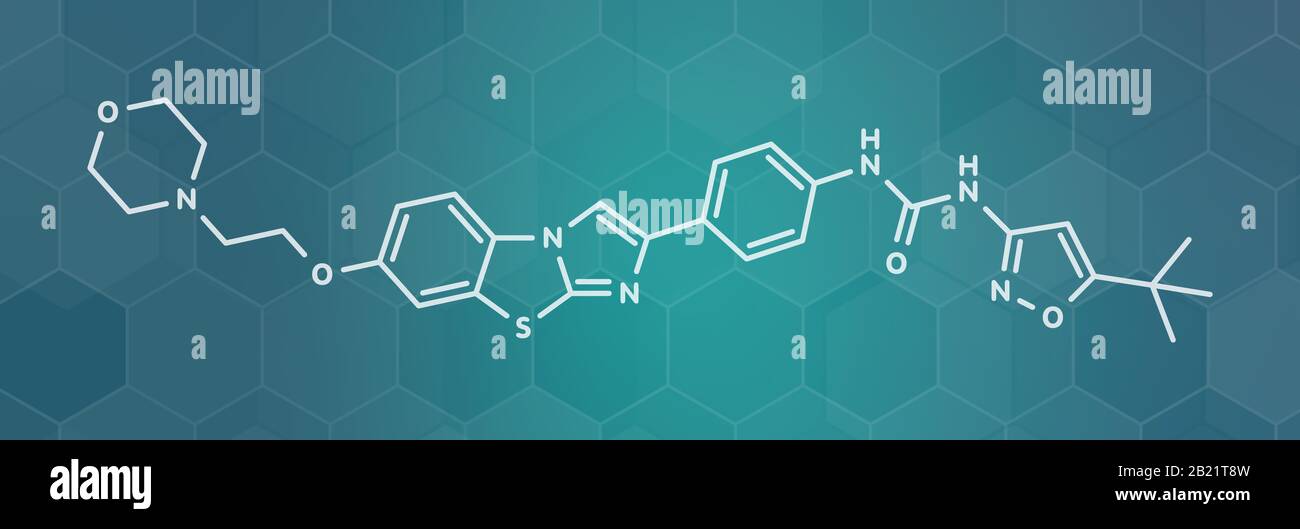
RF2C9JHCK–Quizartinib cancer drug molecule (kinase inhibitor). White skeletal formula on dark teal gradient background with hexagonal pattern.
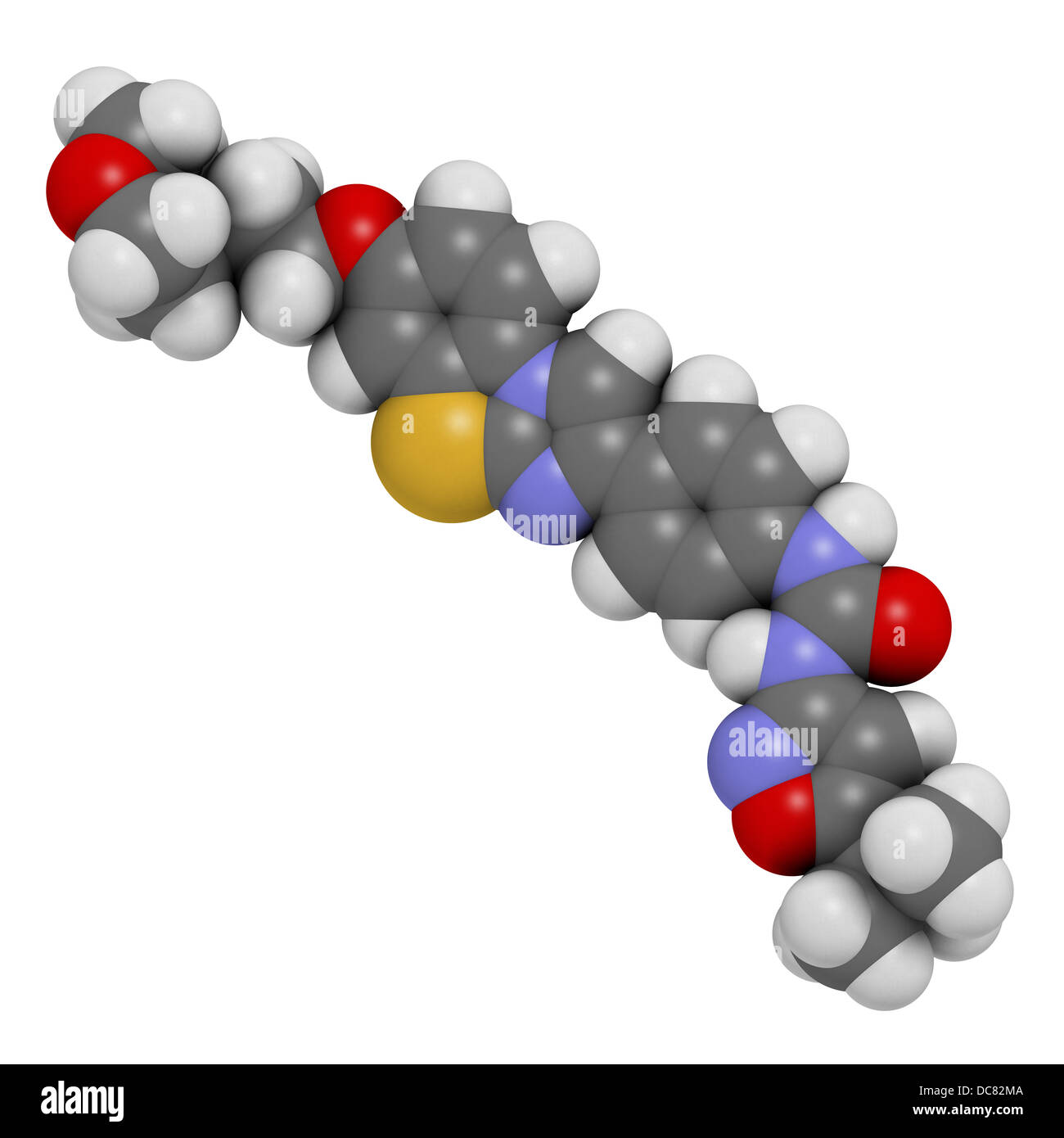
RFDC82MA–Quizartinib investigational acute myeloid leukemia (AML) drug, chemical structure Atoms are represented as spheres.
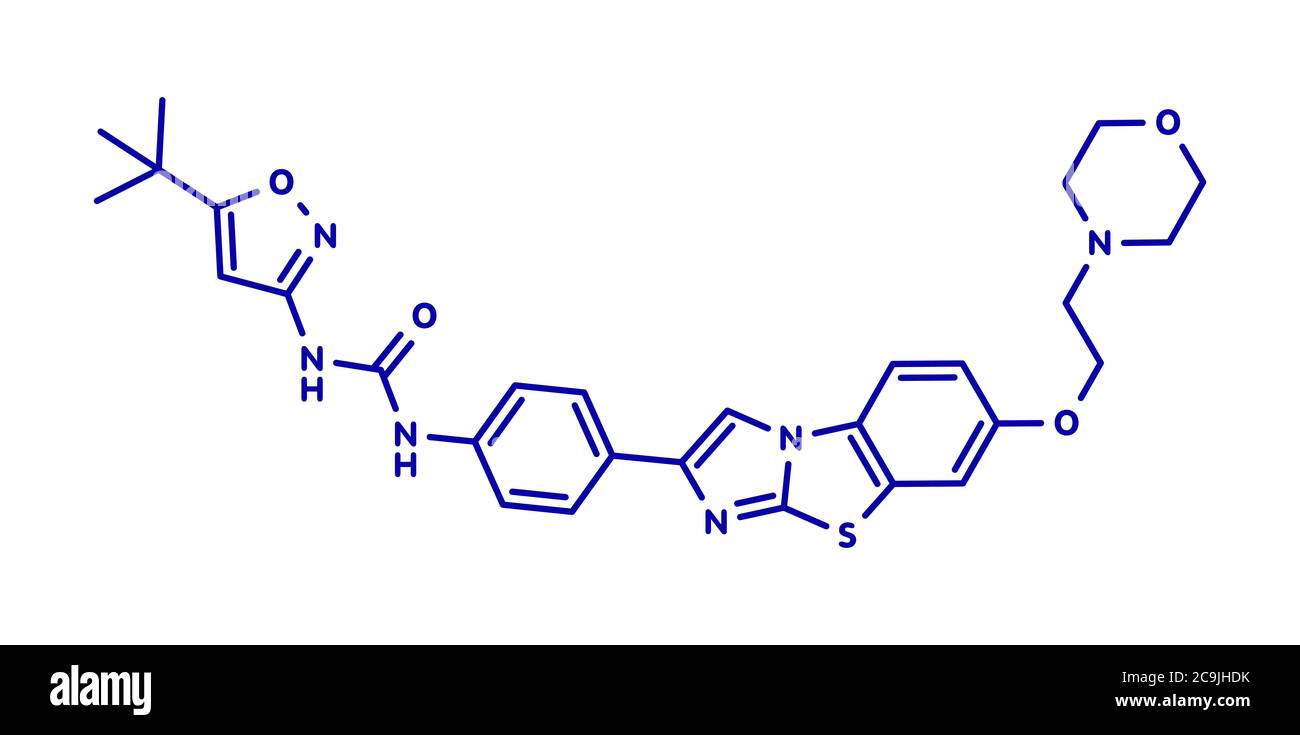
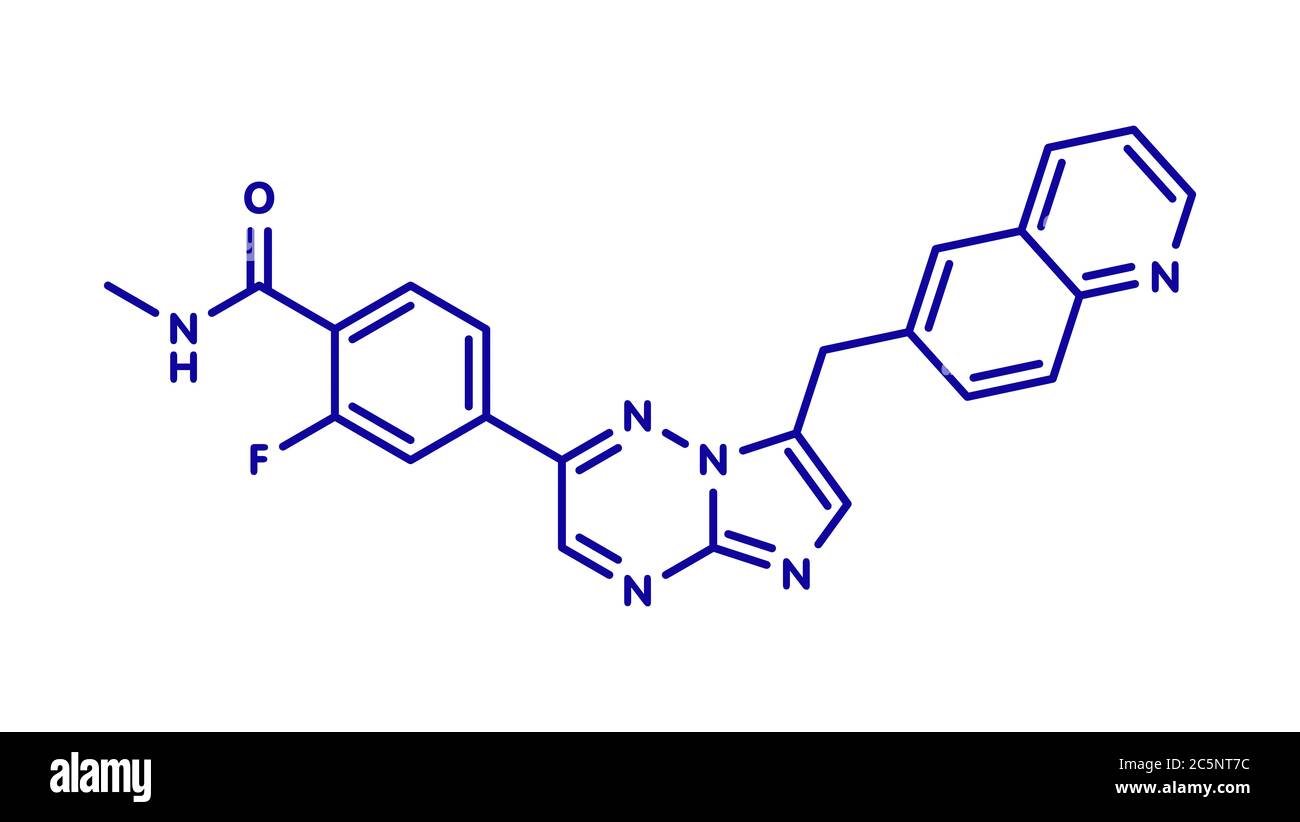
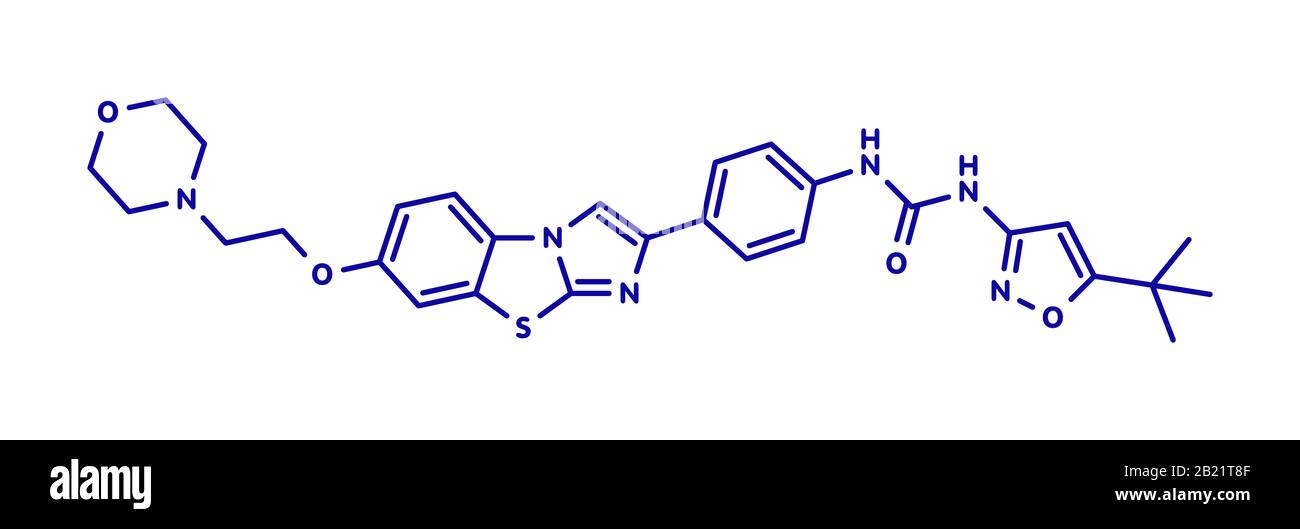
RF2C9JHDK–Quizartinib cancer drug molecule (kinase inhibitor). Blue skeletal formula on white background.
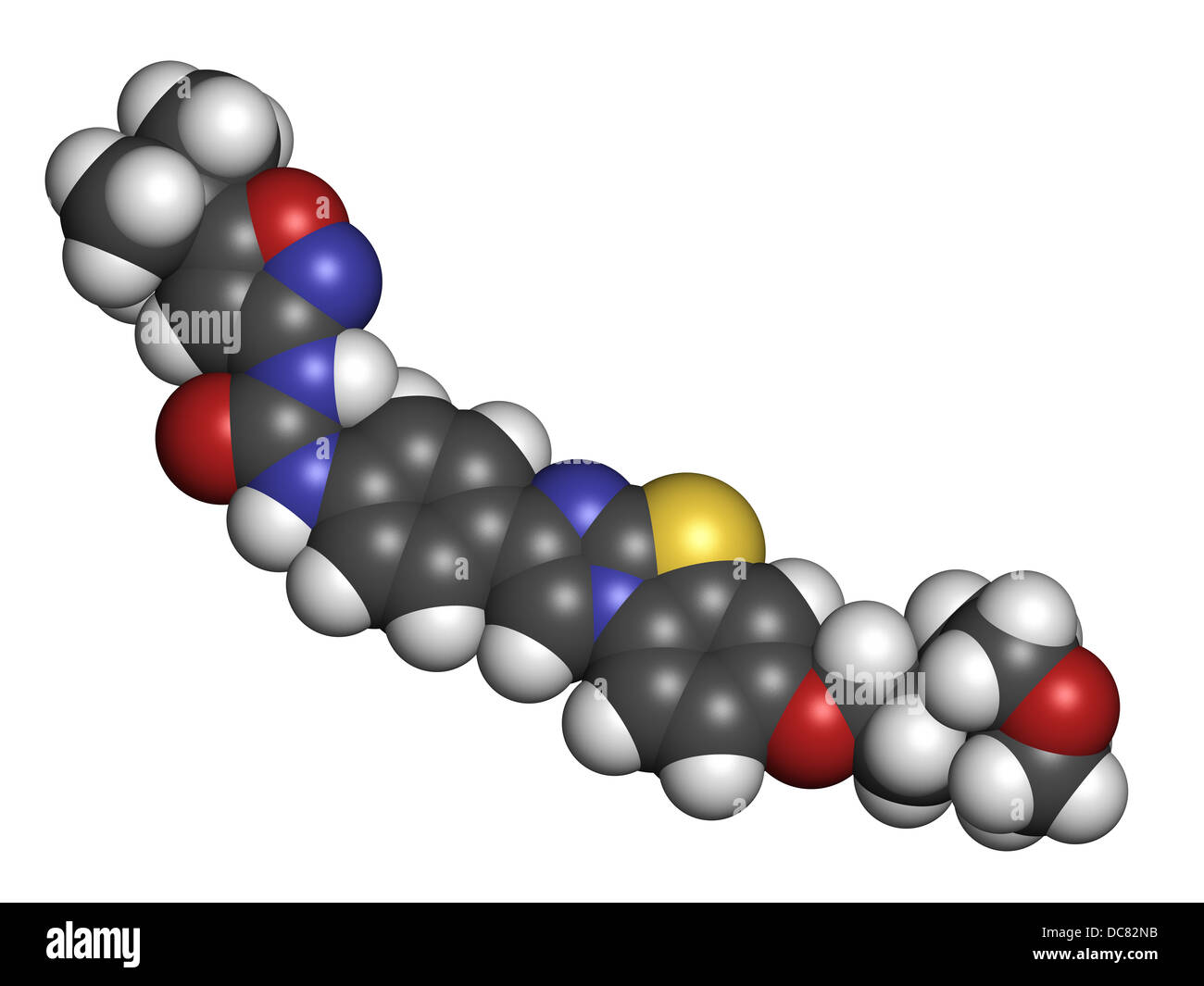
RFDC82NB–Quizartinib investigational acute myeloid leukemia (AML) drug, chemical structure Atoms are represented as spheres.
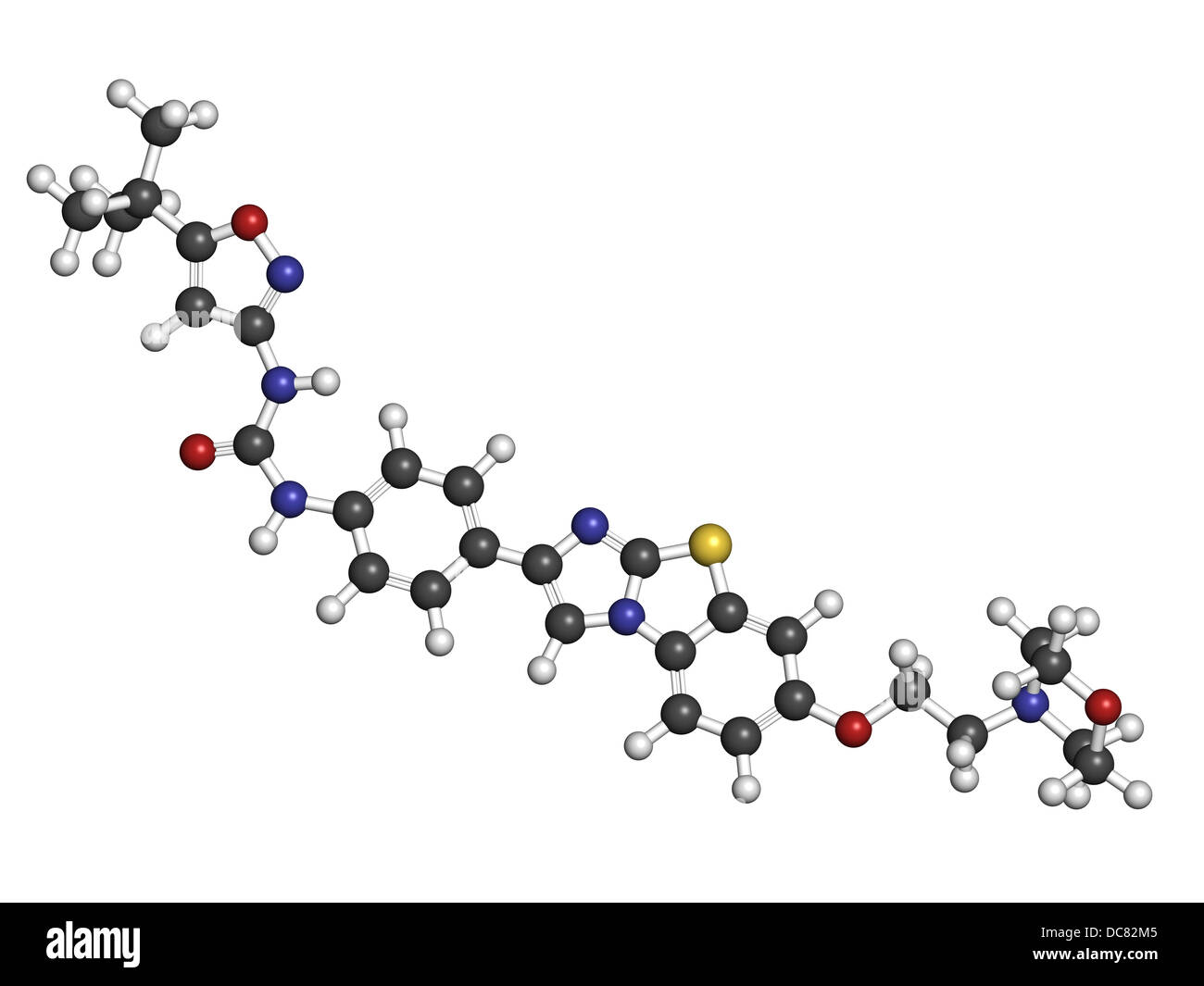
RFDC82M5–Quizartinib investigational acute myeloid leukemia (AML) drug, chemical structure Atoms are represented as spheres.
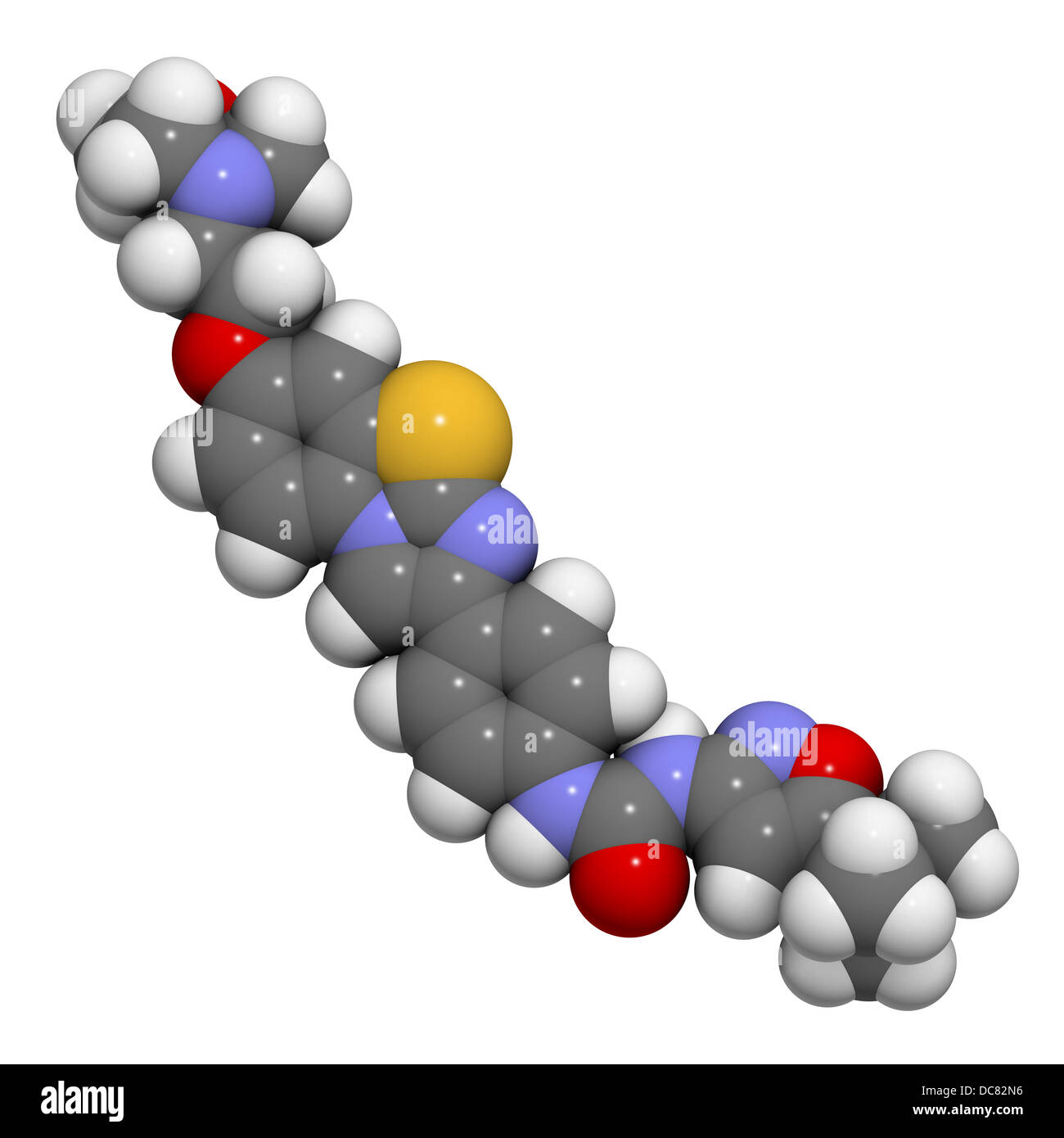
RFDC82N6–Quizartinib investigational acute myeloid leukemia (AML) drug, chemical structure Atoms are represented as spheres.
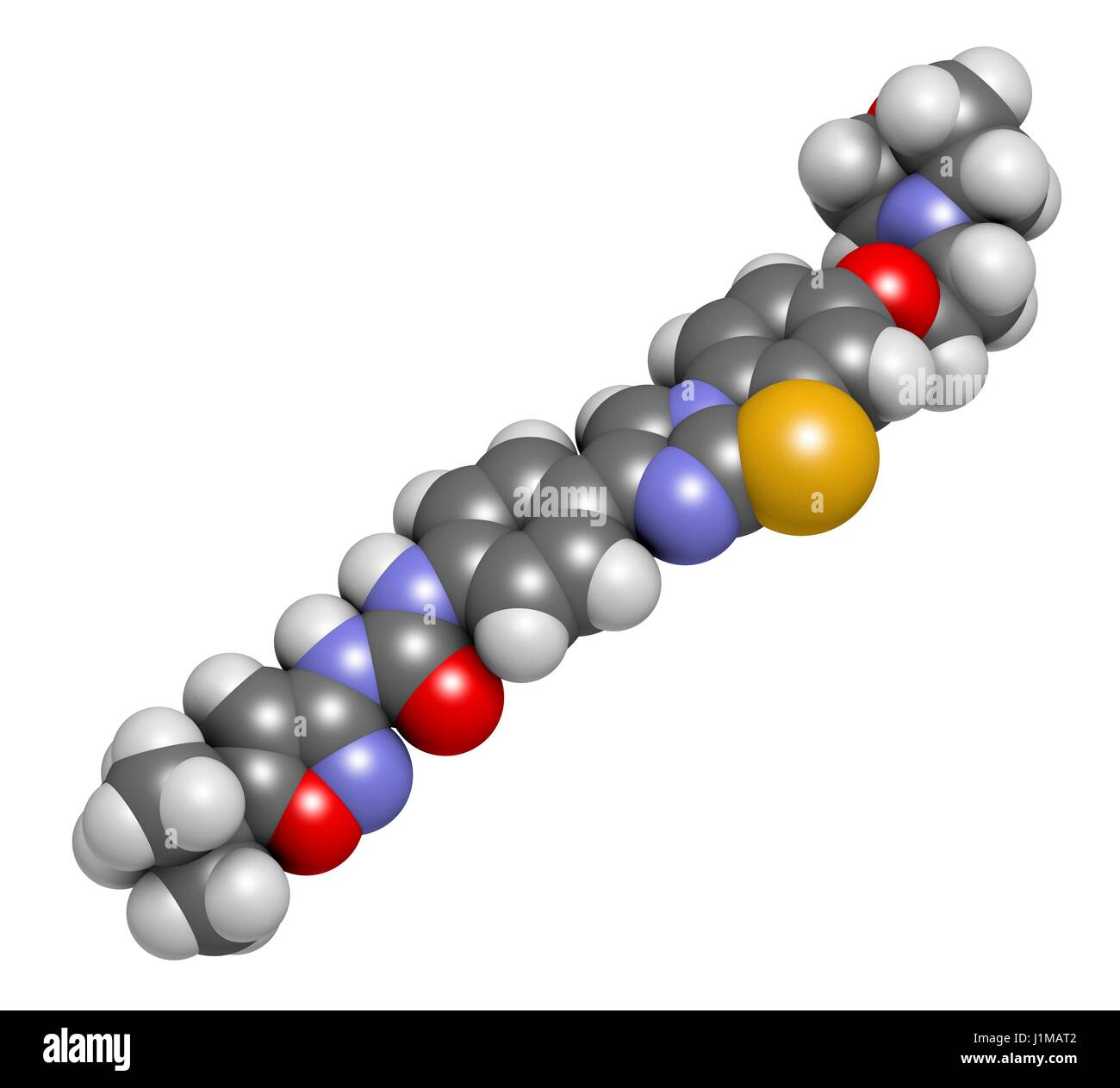
RFJ1MAT2–Quizartinib cancer drug molecule (kinase inhibitor). 3D rendering. Atoms are represented as spheres with conventional colour coding: hydrogen (white), carbon (grey), nitrogen (blue), oxygen (red), sulfur (yellow).
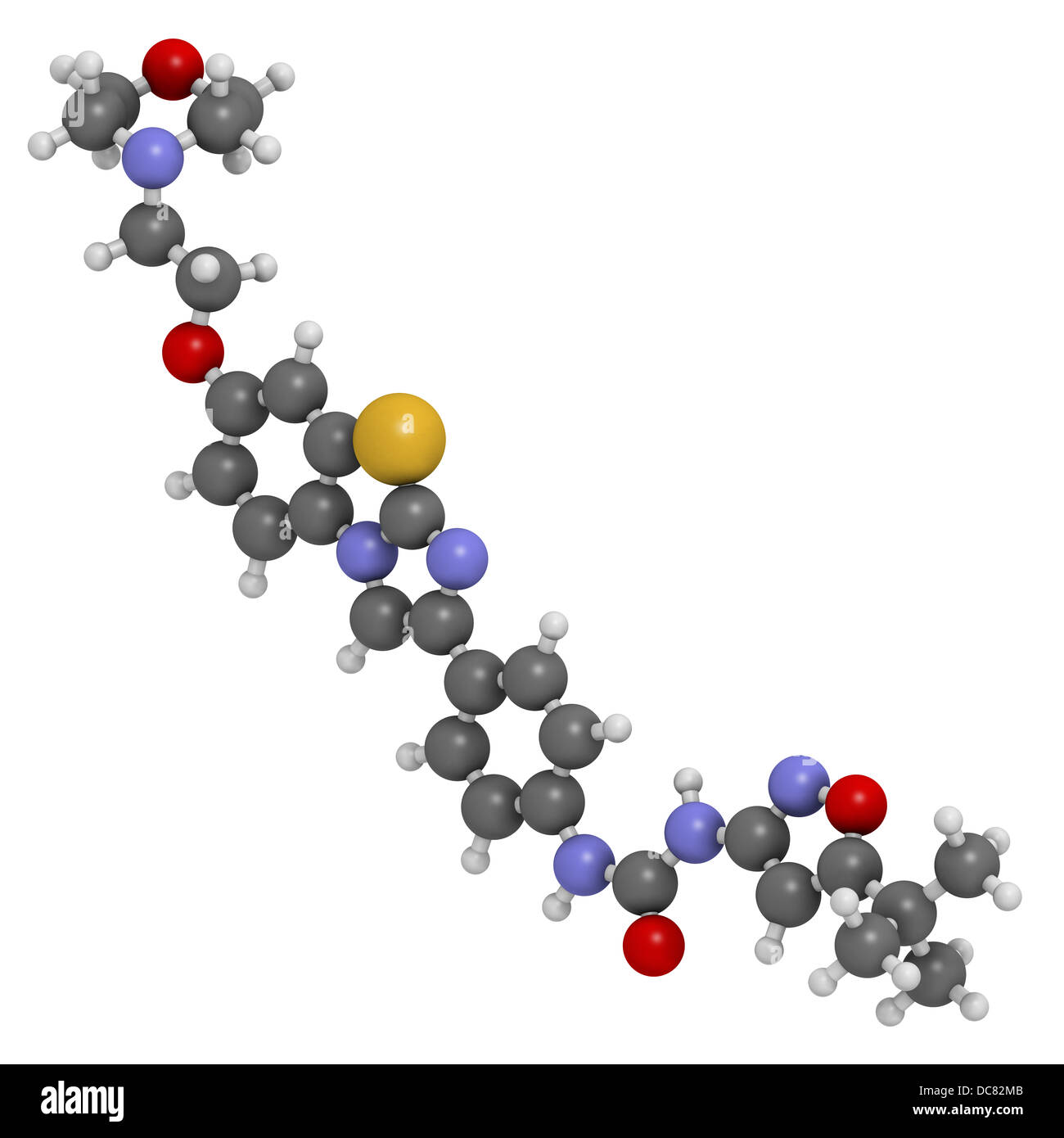
RFDC82MB–Quizartinib investigational acute myeloid leukemia (AML) drug, chemical structure Atoms are represented as spheres.

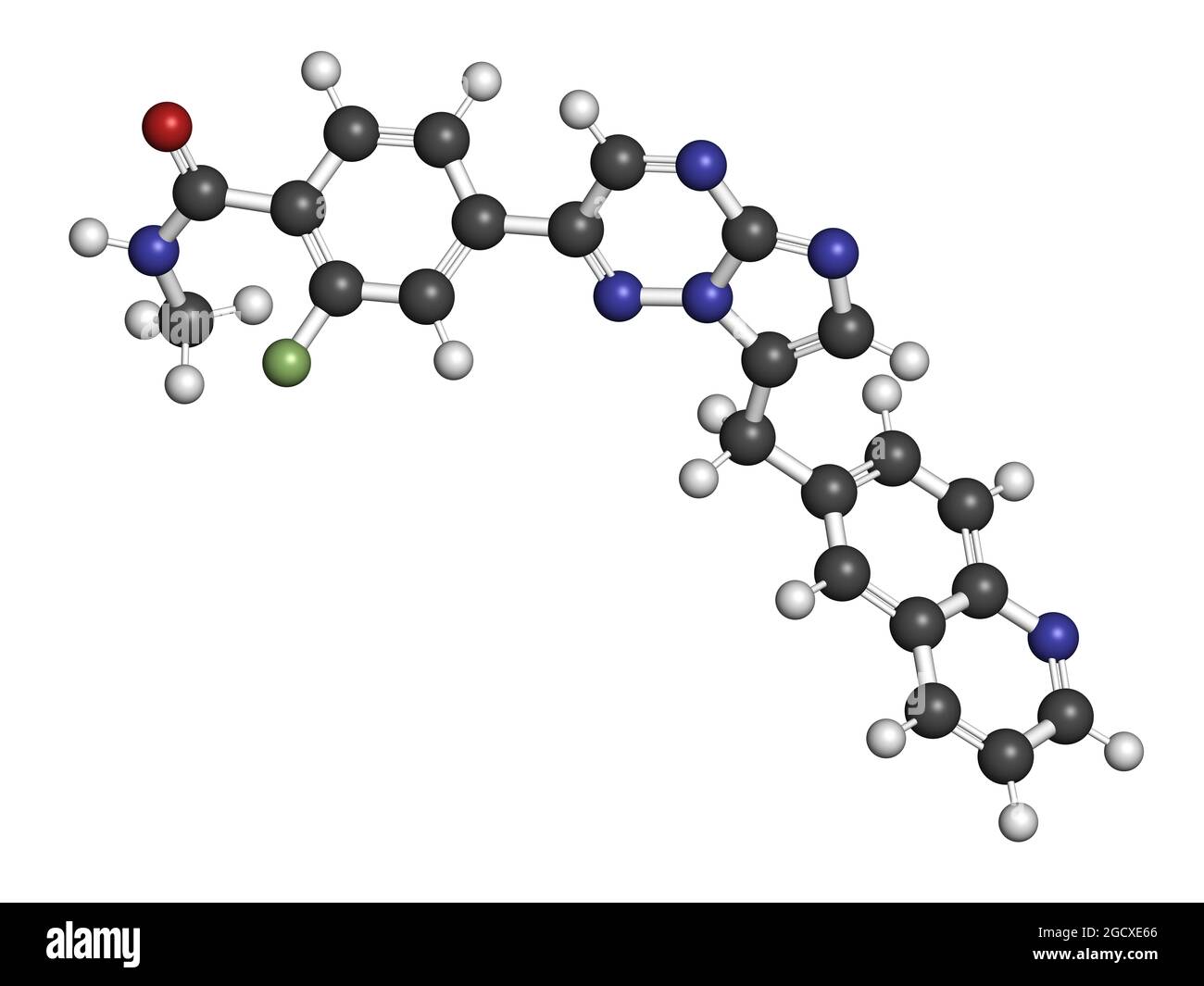
RFJ1MAT3–Quizartinib cancer drug molecule (kinase inhibitor). 3D rendering. Atoms are represented as spheres with conventional colour coding: hydrogen (white), carbon (black), nitrogen (blue), oxygen (red), sulfur (yellow).
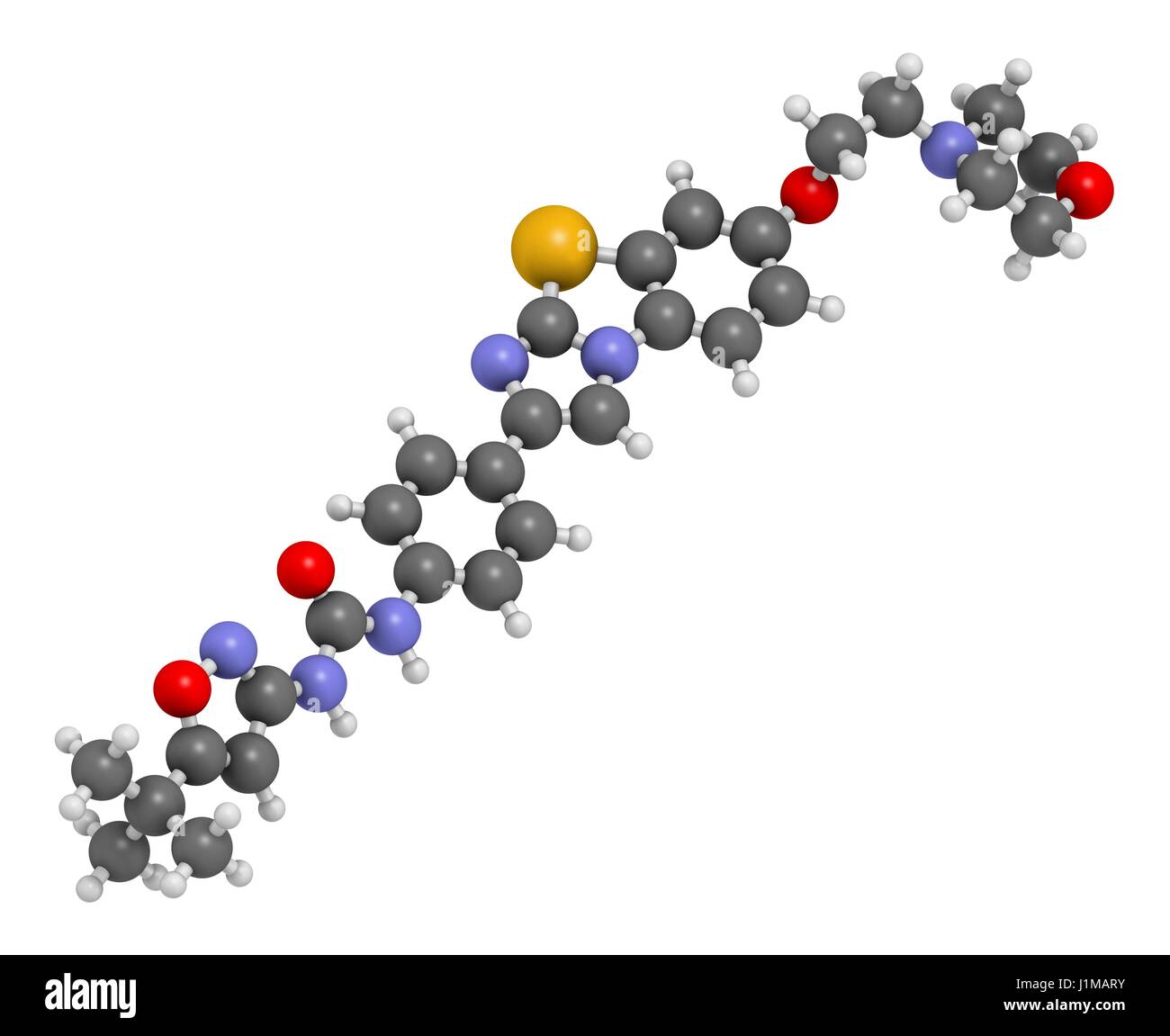
RFJ1MARY–Quizartinib cancer drug molecule (kinase inhibitor). 3D rendering. Atoms are represented as spheres with conventional colour coding: hydrogen (white), carbon (grey), nitrogen (blue), oxygen (red), sulfur (yellow).
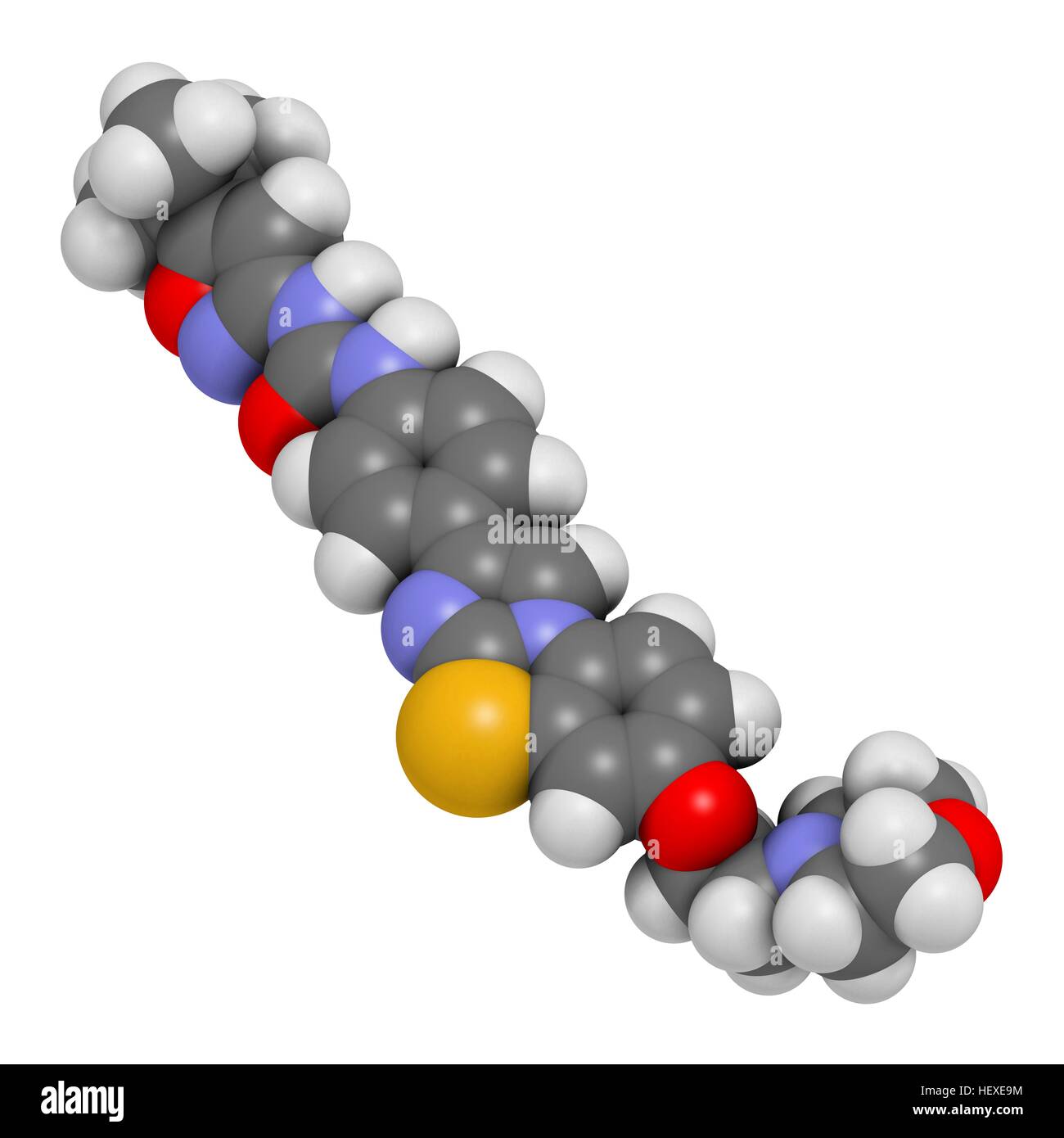
RFHEXE9M–Quizartinib cancer drug molecule (kinase inhibitor), computer illustration. Atoms are represented as spheres with conventional colour coding: hydrogen (white), carbon (grey), nitrogen (blue), oxygen (red), sulphur (yellow).
Search Results for Proto oncogene Stock Photos and Images (62)
Page 1 of 1